Decorate the Examples: A Simple Method of Prompt Design for Biomedical Relation Extraction
Paper and Code
Apr 21, 2022
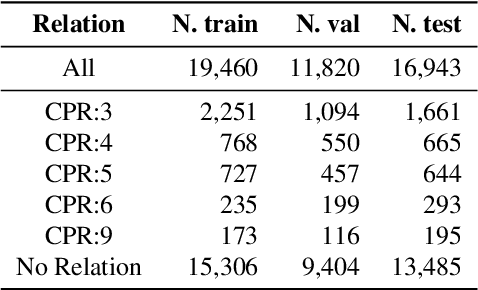
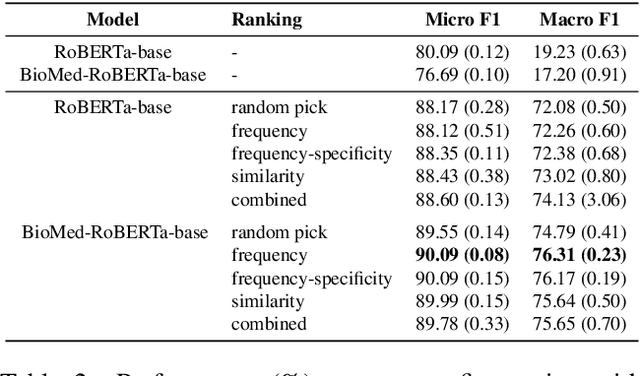
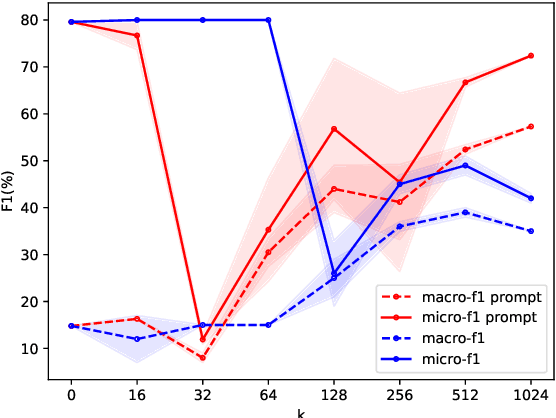
Relation extraction is a core problem for natural language processing in the biomedical domain. Recent research on relation extraction showed that prompt-based learning improves the performance on both fine-tuning on full training set and few-shot training. However, less effort has been made on domain-specific tasks where good prompt design can be even harder. In this paper, we investigate prompting for biomedical relation extraction, with experiments on the ChemProt dataset. We present a simple yet effective method to systematically generate comprehensive prompts that reformulate the relation extraction task as a cloze-test task under a simple prompt formulation. In particular, we experiment with different ranking scores for prompt selection. With BioMed-RoBERTa-base, our results show that prompting-based fine-tuning obtains gains by 14.21 F1 over its regular fine-tuning baseline, and 1.14 F1 over SciFive-Large, the current state-of-the-art on ChemProt. Besides, we find prompt-based learning requires fewer training examples to make reasonable predictions. The results demonstrate the potential of our methods in such a domain-specific relation extraction task.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge