Cross-Organ and Cross-Scanner Adenocarcinoma Segmentation using Rein to Fine-tune Vision Foundation Models
Paper and Code
Sep 19, 2024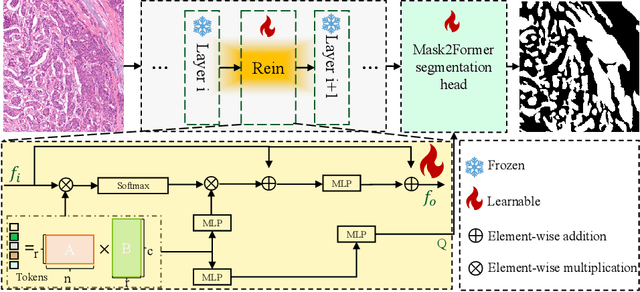
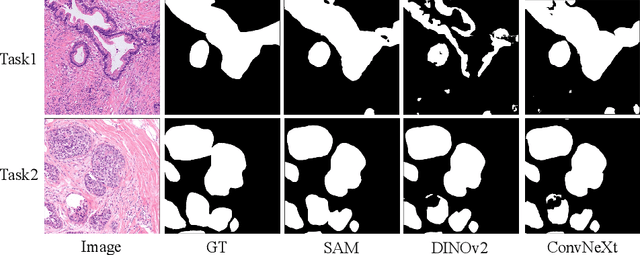

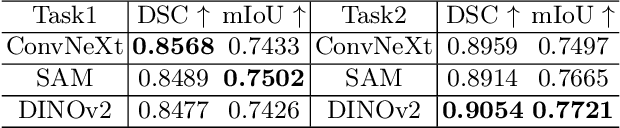

In recent years, significant progress has been made in tumor segmentation within the field of digital pathology. However, variations in organs, tissue preparation methods, and image acquisition processes can lead to domain discrepancies among digital pathology images. To address this problem, in this paper, we use Rein, a fine-tuning method, to parametrically and efficiently fine-tune various vision foundation models (VFMs) for MICCAI 2024 Cross-Organ and Cross-Scanner Adenocarcinoma Segmentation (COSAS2024). The core of Rein consists of a set of learnable tokens, which are directly linked to instances, improving functionality at the instance level in each layer. In the data environment of the COSAS2024 Challenge, extensive experiments demonstrate that Rein fine-tuned the VFMs to achieve satisfactory results. Specifically, we used Rein to fine-tune ConvNeXt and DINOv2. Our team used the former to achieve scores of 0.7719 and 0.7557 on the preliminary test phase and final test phase in task1, respectively, while the latter achieved scores of 0.8848 and 0.8192 on the preliminary test phase and final test phase in task2. Code is available at GitHub.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge