Compositionally Equivariant Representation Learning
Paper and Code
Jun 17, 2023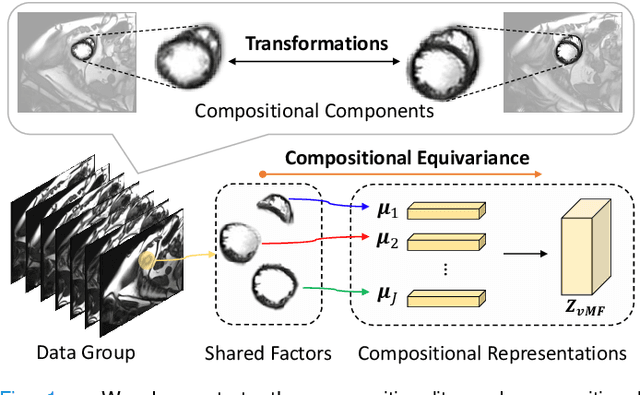
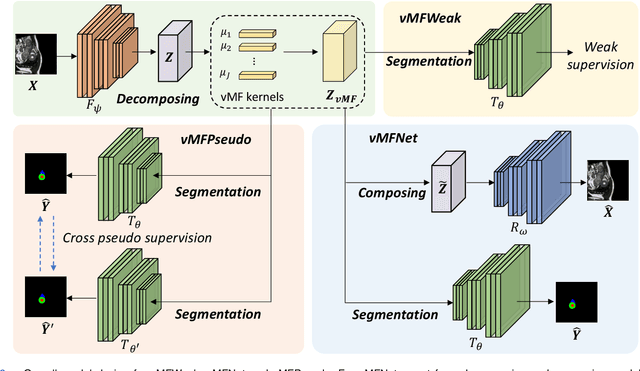
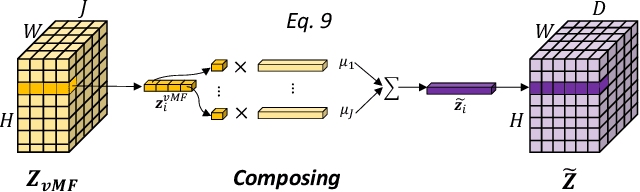
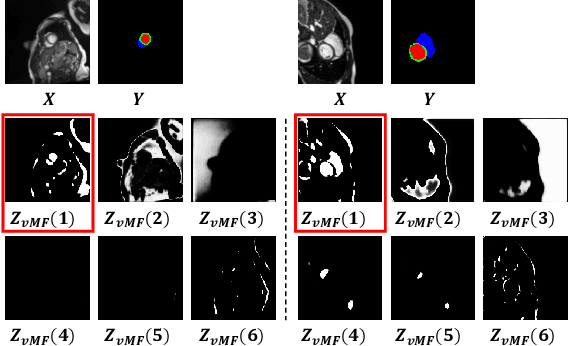
Deep learning models often need sufficient supervision (i.e. labelled data) in order to be trained effectively. By contrast, humans can swiftly learn to identify important anatomy in medical images like MRI and CT scans, with minimal guidance. This recognition capability easily generalises to new images from different medical facilities and to new tasks in different settings. This rapid and generalisable learning ability is largely due to the compositional structure of image patterns in the human brain, which are not well represented in current medical models. In this paper, we study the utilisation of compositionality in learning more interpretable and generalisable representations for medical image segmentation. Overall, we propose that the underlying generative factors that are used to generate the medical images satisfy compositional equivariance property, where each factor is compositional (e.g. corresponds to the structures in human anatomy) and also equivariant to the task. Hence, a good representation that approximates well the ground truth factor has to be compositionally equivariant. By modelling the compositional representations with learnable von-Mises-Fisher (vMF) kernels, we explore how different design and learning biases can be used to enforce the representations to be more compositionally equivariant under un-, weakly-, and semi-supervised settings. Extensive results show that our methods achieve the best performance over several strong baselines on the task of semi-supervised domain-generalised medical image segmentation. Code will be made publicly available upon acceptance at https://github.com/vios-s.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge