Clinical Reading Comprehension: A Thorough Analysis of the emrQA Dataset
Paper and Code
May 01, 2020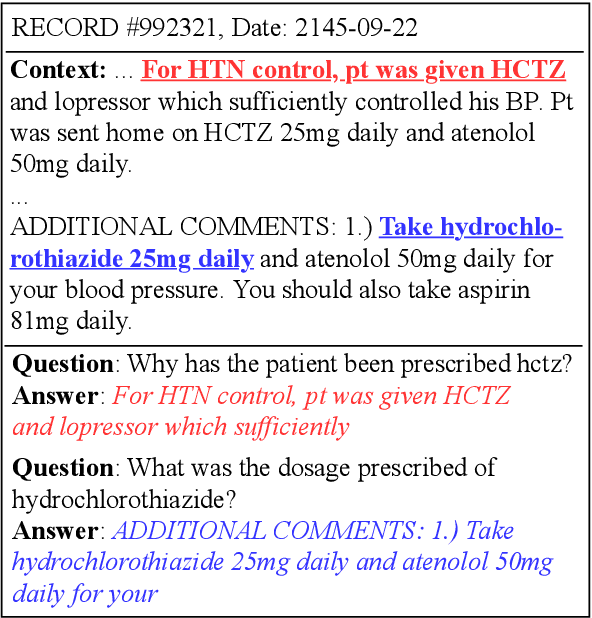


Machine reading comprehension has made great progress in recent years owing to large-scale annotated datasets. In the clinical domain, however, creating such datasets is quite difficult due to the domain expertise required for annotation. Recently, Pampari et al. (EMNLP'18) tackled this issue by using expert-annotated question templates and existing i2b2 annotations to create emrQA, the first large-scale dataset for question answering (QA) based on clinical notes. In this paper, we provide an in-depth analysis of this dataset and the clinical reading comprehension (CliniRC) task. From our qualitative analysis, we find that (i) emrQA answers are often incomplete, and (ii) emrQA questions are often answerable without using domain knowledge. From our quantitative experiments, surprising results include that (iii) using a small sampled subset (5%-20%), we can obtain roughly equal performance compared to the model trained on the entire dataset, (iv) this performance is close to human expert's performance, and (v) BERT models do not beat the best performing base model. Following our analysis of the emrQA, we further explore two desired aspects of CliniRC systems: the ability to utilize clinical domain knowledge and to generalize to unseen questions and contexts. We argue that both should be considered when creating future datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge