Beyond Voxel Prediction Uncertainty: Identifying brain lesions you can trust
Paper and Code
Sep 22, 2022
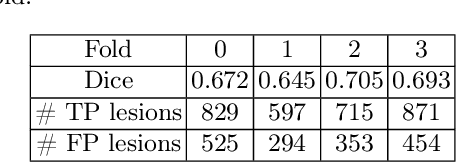
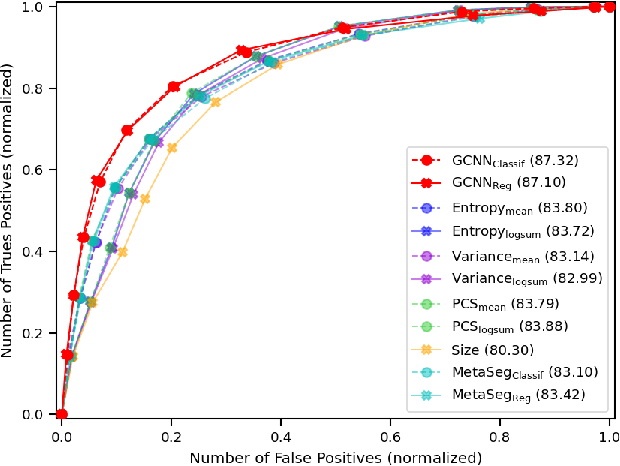
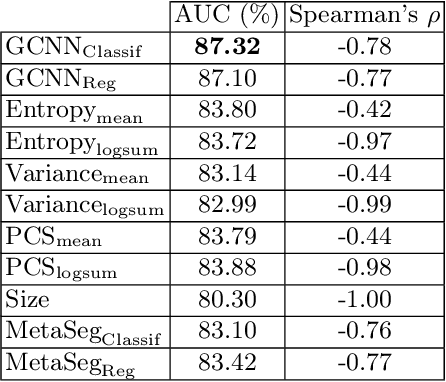
Deep neural networks have become the gold-standard approach for the automated segmentation of 3D medical images. Their full acceptance by clinicians remains however hampered by the lack of intelligible uncertainty assessment of the provided results. Most approaches to quantify their uncertainty, such as the popular Monte Carlo dropout, restrict to some measure of uncertainty in prediction at the voxel level. In addition not to be clearly related to genuine medical uncertainty, this is not clinically satisfying as most objects of interest (e.g. brain lesions) are made of groups of voxels whose overall relevance may not simply reduce to the sum or mean of their individual uncertainties. In this work, we propose to go beyond voxel-wise assessment using an innovative Graph Neural Network approach, trained from the outputs of a Monte Carlo dropout model. This network allows the fusion of three estimators of voxel uncertainty: entropy, variance, and model's confidence; and can be applied to any lesion, regardless of its shape or size. We demonstrate the superiority of our approach for uncertainty estimate on a task of Multiple Sclerosis lesions segmentation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge