Analyzing Regional Organization of the Human Hippocampus in 3D-PLI Using Contrastive Learning and Geometric Unfolding
Paper and Code
Feb 27, 2024
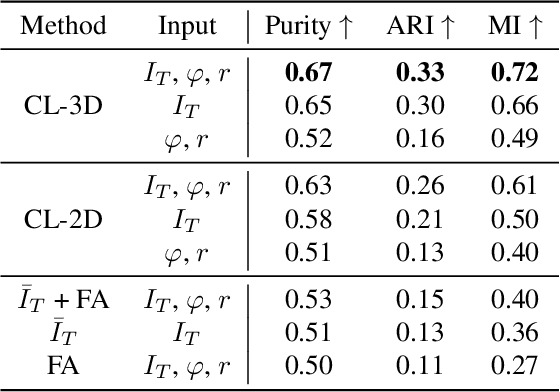
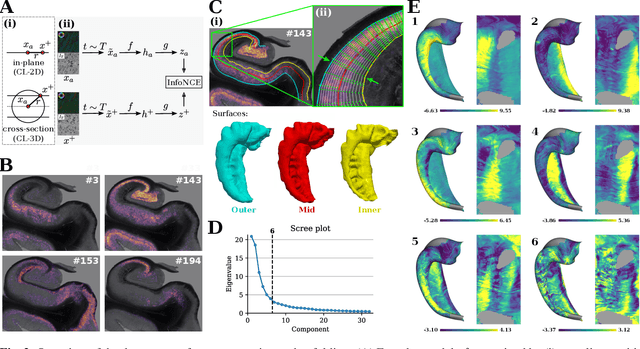
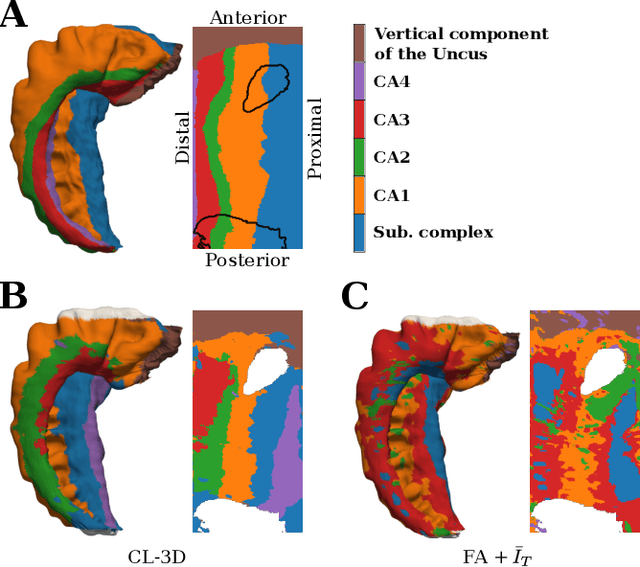
Understanding the cortical organization of the human brain requires interpretable descriptors for distinct structural and functional imaging data. 3D polarized light imaging (3D-PLI) is an imaging modality for visualizing fiber architecture in postmortem brains with high resolution that also captures the presence of cell bodies, for example, to identify hippocampal subfields. The rich texture in 3D-PLI images, however, makes this modality particularly difficult to analyze and best practices for characterizing architectonic patterns still need to be established. In this work, we demonstrate a novel method to analyze the regional organization of the human hippocampus in 3D-PLI by combining recent advances in unfolding methods with deep texture features obtained using a self-supervised contrastive learning approach. We identify clusters in the representations that correspond well with classical descriptions of hippocampal subfields, lending validity to the developed methodology.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge