A Dual-branch Self-supervised Representation Learning Framework for Tumour Segmentation in Whole Slide Images
Paper and Code
Mar 20, 2023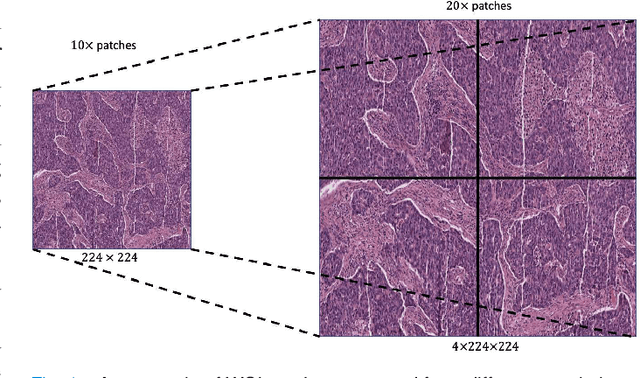
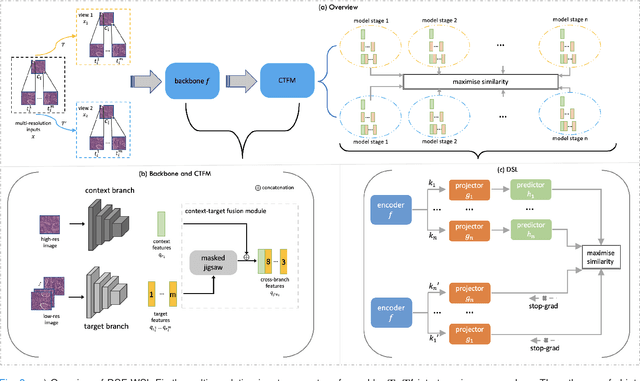
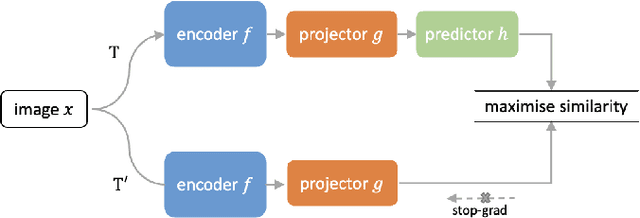
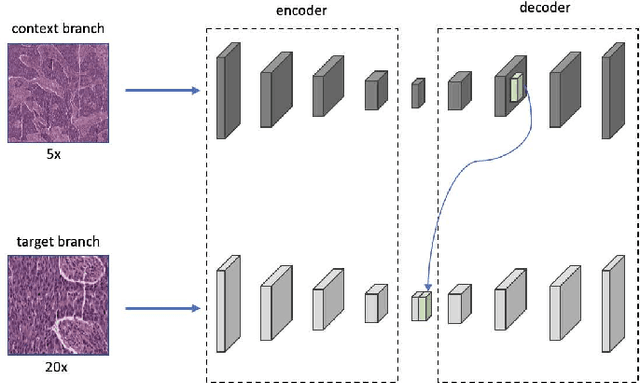
Supervised deep learning methods have achieved considerable success in medical image analysis, owing to the availability of large-scale and well-annotated datasets. However, creating such datasets for whole slide images (WSIs) in histopathology is a challenging task due to their gigapixel size. In recent years, self-supervised learning (SSL) has emerged as an alternative solution to reduce the annotation overheads in WSIs, as it does not require labels for training. These SSL approaches, however, are not designed for handling multi-resolution WSIs, which limits their performance in learning discriminative image features. In this paper, we propose a Dual-branch SSL Framework for WSI tumour segmentation (DSF-WSI) that can effectively learn image features from multi-resolution WSIs. Our DSF-WSI connected two branches and jointly learnt low and high resolution WSIs in a self-supervised manner. Moreover, we introduced a novel Context-Target Fusion Module (CTFM) and a masked jigsaw pretext task to align the learnt multi-resolution features. Furthermore, we designed a Dense SimSiam Learning (DSL) strategy to maximise the similarity of different views of WSIs, enabling the learnt representations to be more efficient and discriminative. We evaluated our method using two public datasets on breast and liver cancer segmentation tasks. The experiment results demonstrated that our DSF-WSI can effectively extract robust and efficient representations, which we validated through subsequent fine-tuning and semi-supervised settings. Our proposed method achieved better accuracy than other state-of-the-art approaches. Code is available at https://github.com/Dylan-H-Wang/dsf-wsi.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge