Yen-Chu Lin
Toward Drug-Target Interaction Prediction via Ensemble Modeling and Transfer Learning
Jul 28, 2021
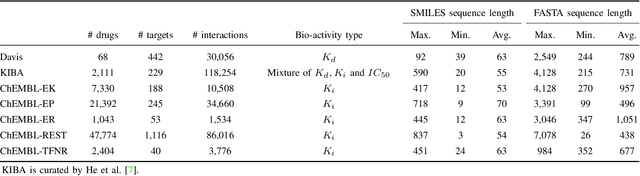
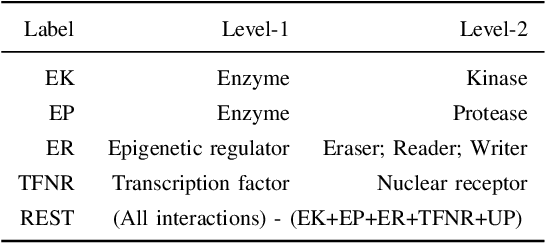

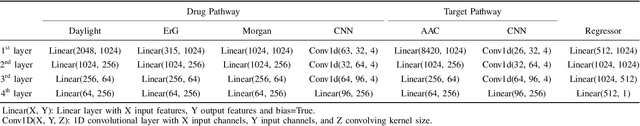
Abstract:Drug-target interaction (DTI) prediction plays a crucial role in drug discovery, and deep learning approaches have achieved state-of-the-art performance in this field. We introduce an ensemble of deep learning models (EnsembleDLM) for DTI prediction. EnsembleDLM only uses the sequence information of chemical compounds and proteins, and it aggregates the predictions from multiple deep neural networks. This approach not only achieves state-of-the-art performance in Davis and KIBA datasets but also reaches cutting-edge performance in the cross-domain applications across different bio-activity types and different protein classes. We also demonstrate that EnsembleDLM achieves a good performance (Pearson correlation coefficient and concordance index > 0.8) in the new domain with approximately 50% transfer learning data, i.e., the training set has twice as much data as the test set.
Combating small molecule aggregation with machine learning
May 01, 2021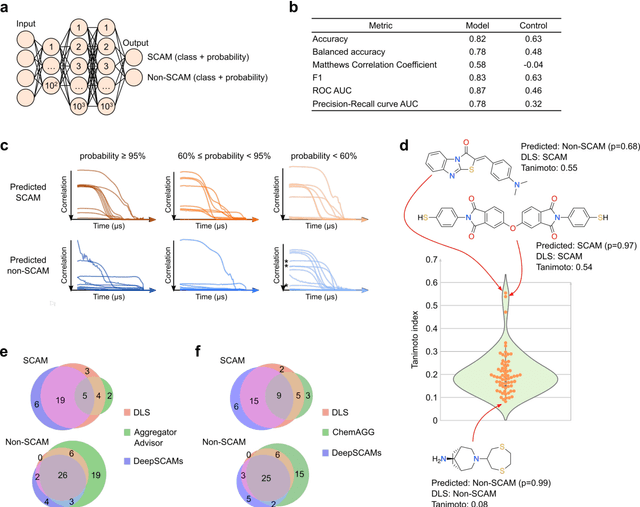
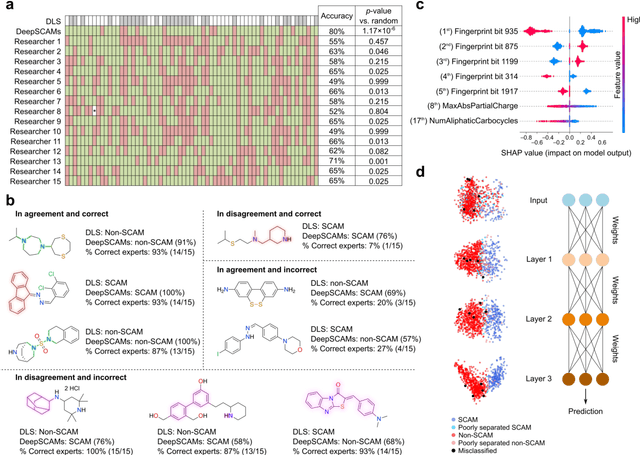
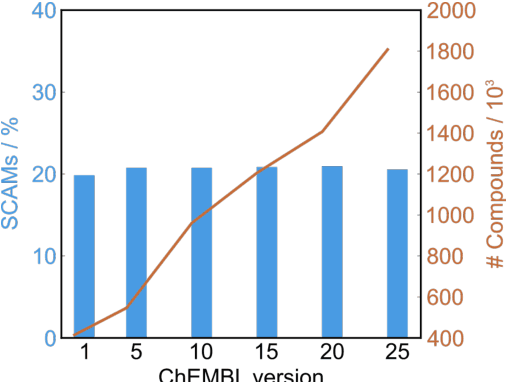
Abstract:Biological screens are plagued by false positive hits resulting from aggregation. Thus, methods to triage small colloidally aggregating molecules (SCAMs) are in high demand. Herein, we disclose a bespoke machine-learning tool to confidently and intelligibly flag such entities. Our data demonstrate an unprecedented utility of machine learning for predicting SCAMs, achieving 80% of correct predictions in a challenging out-of-sample validation. The tool outperformed a panel of expert chemists, who correctly predicted 61 +/- 7% of the same test molecules in a Turing-like test. Further, the computational routine provided insight into molecular features governing aggregation that had remained hidden to expert intuition. Leveraging our tool, we quantify that up to 15-20% of ligands in publicly available chemogenomic databases have the high potential to aggregate at typical screening concentrations, imposing caution in systems biology and drug design programs. Our approach provides a means to augment human intuition, mitigate attrition and a pathway to accelerate future molecular medicine.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge