Shuangge Ma
Sparse Bayesian Deep Functional Learning with Structured Region Selection
Feb 24, 2026Abstract:In modern applications such as ECG monitoring, neuroimaging, wearable sensing, and industrial equipment diagnostics, complex and continuously structured data are ubiquitous, presenting both challenges and opportunities for functional data analysis. However, existing methods face a critical trade-off: conventional functional models are limited by linearity, whereas deep learning approaches lack interpretable region selection for sparse effects. To bridge these gaps, we propose a sparse Bayesian functional deep neural network (sBayFDNN). It learns adaptive functional embeddings through a deep Bayesian architecture to capture complex nonlinear relationships, while a structured prior enables interpretable, region-wise selection of influential domains with quantified uncertainty. Theoretically, we establish rigorous approximation error bounds, posterior consistency, and region selection consistency. These results provide the first theoretical guarantees for a Bayesian deep functional model, ensuring its reliability and statistical rigor. Empirically, comprehensive simulations and real-world studies confirm the effectiveness and superiority of sBayFDNN. Crucially, sBayFDNN excels in recognizing intricate dependencies for accurate predictions and more precisely identifies functionally meaningful regions, capabilities fundamentally beyond existing approaches.
Penalized Generative Variable Selection
Feb 26, 2024Abstract:Deep networks are increasingly applied to a wide variety of data, including data with high-dimensional predictors. In such analysis, variable selection can be needed along with estimation/model building. Many of the existing deep network studies that incorporate variable selection have been limited to methodological and numerical developments. In this study, we consider modeling/estimation using the conditional Wasserstein Generative Adversarial networks. Group Lasso penalization is applied for variable selection, which may improve model estimation/prediction, interpretability, stability, etc. Significantly advancing from the existing literature, the analysis of censored survival data is also considered. We establish the convergence rate for variable selection while considering the approximation error, and obtain a more efficient distribution estimation. Simulations and the analysis of real experimental data demonstrate satisfactory practical utility of the proposed analysis.
Structural modeling using overlapped group penalties for discovering predictive biomarkers for subgroup analysis
Apr 26, 2019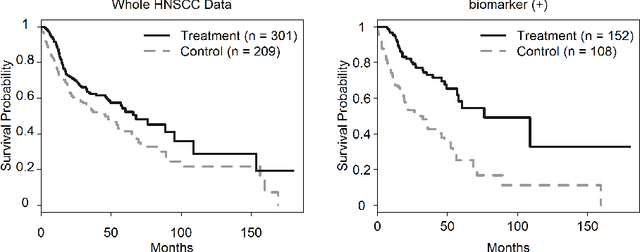

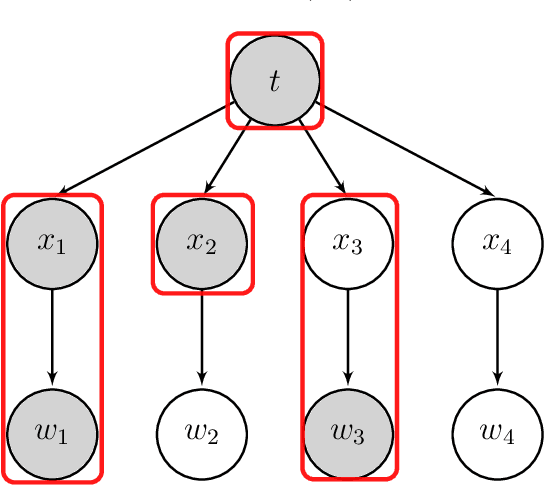

Abstract:The identification of predictive biomarkers from a large scale of covariates for subgroup analysis has attracted fundamental attention in medical research. In this article, we propose a generalized penalized regression method with a novel penalty function, for enforcing the hierarchy structure between the prognostic and predictive effects, such that a nonzero predictive effect must induce its ancestor prognostic effects being nonzero in the model. Our method is able to select useful predictive biomarkers by yielding a sparse, interpretable, and predictable model for subgroup analysis, and can deal with different types of response variable such as continuous, categorical, and time-to-event data. We show that our method is asymptotically consistent under some regularized conditions. To minimize the generalized penalized regression model, we propose a novel integrative optimization algorithm by integrating the majorization-minimization and the alternating direction method of multipliers, which is named after \texttt{smog}. The enriched simulation study and real case study demonstrate that our method is very powerful for discovering the true predictive biomarkers and identifying subgroups of patients.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge