Shahzad Mirza
Machine learning techniques to identify antibiotic resistance in patients diagnosed with various skin and soft tissue infections
Feb 28, 2022
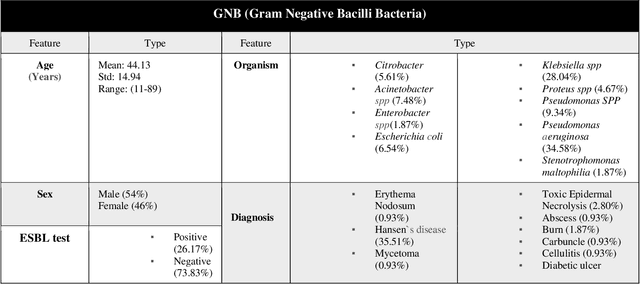
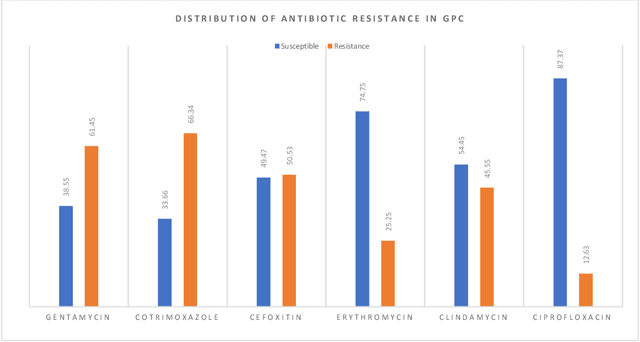

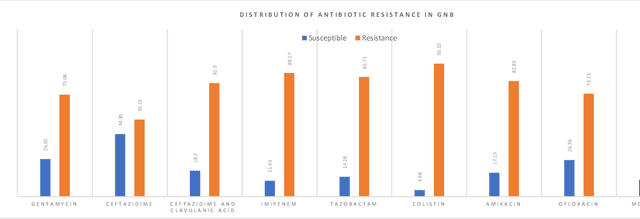
Abstract:Skin and soft tissue infections (SSTIs) are among the most frequently observed diseases in ambulatory and hospital settings. Resistance of diverse bacterial pathogens to antibiotics is a significant cause of severe SSTIs, and treatment failure results in morbidity, mortality, and increased cost of hospitalization. Therefore, antimicrobial surveillance is essential to predict antibiotic resistance trends and monitor the results of medical interventions. To address this, we developed machine learning (ML) models (deep and conventional algorithms) to predict antimicrobial resistance using antibiotic susceptibility testing (ABST) data collected from patients clinically diagnosed with primary and secondary pyoderma over a period of one year. We trained an individual ML algorithm on each antimicrobial family to determine whether a Gram-Positive Cocci (GPC) or Gram-Negative Bacilli (GNB) bacteria will resist the corresponding antibiotic. For this purpose, clinical and demographic features from the patient and data from ABST were employed in training. We achieved an Area Under the Curve (AUC) of 0.68-0.98 in GPC and 0.56-0.93 in GNB bacteria, depending on the antimicrobial family. We also conducted a correlation analysis to determine the linear relationship between each feature and antimicrobial families in different bacteria. ML techniques suggest that a predictable nonlinear relationship exists between patients' clinical-demographic characteristics and antibiotic resistance; however, the accuracy of this prediction depends on the type of the antimicrobial family.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge