Ruggiero Santeramo
MamaDino: A Hybrid Vision Model for Breast Cancer 3-Year Risk Prediction
Feb 14, 2026Abstract:Breast cancer screening programmes increasingly seek to move from one-size-fits-all interval to risk-adapted and personalized strategies. Deep learning (DL) has enabled image-based risk models with stronger 1- to 5-year prediction than traditional clinical models, but leading systems (e.g., Mirai) typically use convolutional backbones, very high-resolution inputs (>1M pixels) and simple multi-view fusion, with limited explicit modelling of contralateral asymmetry. We hypothesised that combining complementary inductive biases (convolutional and transformer-based) with explicit contralateral asymmetry modelling would allow us to match state-of-the-art 3-year risk prediction performance even when operating on substantially lower-resolution mammograms, indicating that using less detailed images in a more structured way can recover state-of-the-art accuracy. We present MamaDino, a mammography-aware multi-view attentional DINO model. MamaDino fuses frozen self-supervised DINOv3 ViT-S features with a trainable CNN encoder at 512x512 resolution, and aggregates bilateral breast information via a BilateralMixer to output a 3-year breast cancer risk score. We train on 53,883 women from OPTIMAM (UK) and evaluate on matched 3-year case-control cohorts: an in-distribution test set from four screening sites and an external out-of-distribution cohort from an unseen site. At breast-level, MamaDino matches Mirai on both internal and external tests while using ~13x fewer input pixels. Adding the BilateralMixer improves discrimination to AUC 0.736 (vs 0.713) in-distribution and 0.677 (vs 0.666) out-of-distribution, with consistent performance across age, ethnicity, scanner, tumour type and grade. These findings demonstrate that explicit contralateral modelling and complementary inductive biases enable predictions that match Mirai, despite operating on substantially lower-resolution mammograms.
Longitudinal detection of radiological abnormalities with time-modulated LSTM
Jul 16, 2018
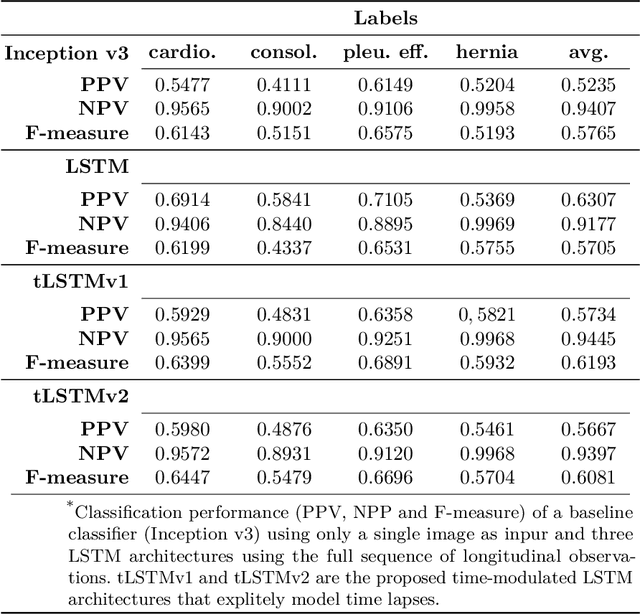

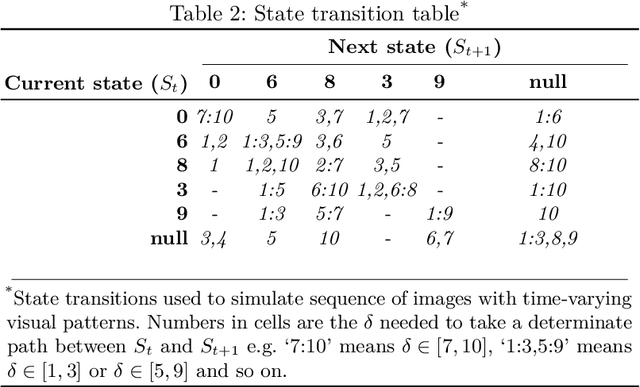
Abstract:Convolutional neural networks (CNNs) have been successfully employed in recent years for the detection of radiological abnormalities in medical images such as plain x-rays. To date, most studies use CNNs on individual examinations in isolation and discard previously available clinical information. In this study we set out to explore whether Long-Short-Term-Memory networks (LSTMs) can be used to improve classification performance when modelling the entire sequence of radiographs that may be available for a given patient, including their reports. A limitation of traditional LSTMs, though, is that they implicitly assume equally-spaced observations, whereas the radiological exams are event-based, and therefore irregularly sampled. Using both a simulated dataset and a large-scale chest x-ray dataset, we demonstrate that a simple modification of the LSTM architecture, which explicitly takes into account the time lag between consecutive observations, can boost classification performance. Our empirical results demonstrate improved detection of commonly reported abnormalities on chest x-rays such as cardiomegaly, consolidation, pleural effusion and hiatus hernia.
* Submitted to 4th MICCAI Workshop on Deep Learning in Medical Imaging Analysis
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge