Paul C. Blainey
Pseudo-Labeling Enhanced by Privileged Information and Its Application to In Situ Sequencing Images
Jun 28, 2023



Abstract:Various strategies for label-scarce object detection have been explored by the computer vision research community. These strategies mainly rely on assumptions that are specific to natural images and not directly applicable to the biological and biomedical vision domains. For example, most semi-supervised learning strategies rely on a small set of labeled data as a confident source of ground truth. In many biological vision applications, however, the ground truth is unknown and indirect information might be available in the form of noisy estimations or orthogonal evidence. In this work, we frame a crucial problem in spatial transcriptomics - decoding barcodes from In-Situ-Sequencing (ISS) images - as a semi-supervised object detection (SSOD) problem. Our proposed framework incorporates additional available sources of information into a semi-supervised learning framework in the form of privileged information. The privileged information is incorporated into the teacher's pseudo-labeling in a teacher-student self-training iteration. Although the available privileged information could be data domain specific, we have introduced a general strategy of pseudo-labeling enhanced by privileged information (PLePI) and exemplified the concept using ISS images, as well on the COCO benchmark using extra evidence provided by CLIP.
* This paper has been accepted for publication at IJCAI 2023
A Bayesian model of acquisition and clearance of bacterial colonization
Nov 27, 2018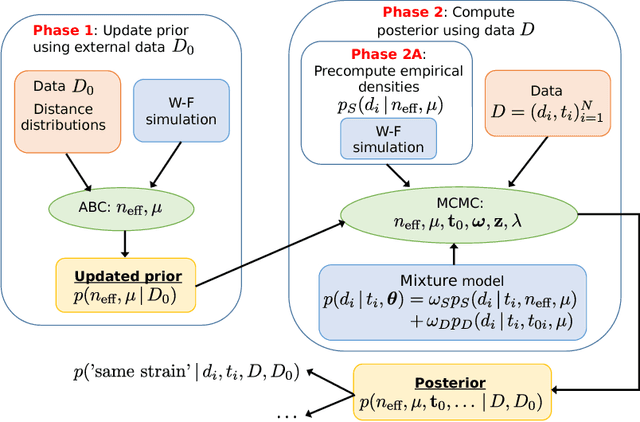
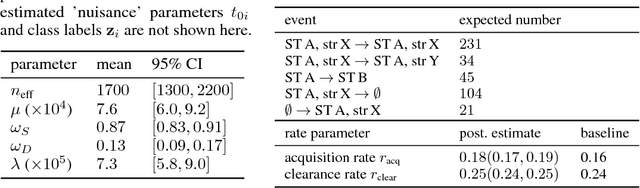



Abstract:Bacterial populations that colonize a host play important roles in host health, including serving as a reservoir that transmits to other hosts and from which invasive strains emerge, thus emphasizing the importance of understanding rates of acquisition and clearance of colonizing populations. Studies of colonization dynamics have been based on assessment of whether serial samples represent a single population or distinct colonization events. A common solution to estimate acquisition and clearance rates is to use a fixed genetic distance threshold. However, this approach is often inadequate to account for the diversity of the underlying within-host evolving population, the time intervals between consecutive measurements, and the uncertainty in the estimated acquisition and clearance rates. Here, we summarize recently submitted work \cite{jarvenpaa2018named} and present a Bayesian model that provides probabilities of whether two strains should be considered the same, allowing to determine bacterial clearance and acquisition from genomes sampled over time. We explicitly model the within-host variation using population genetic simulation, and the inference is done by combining information from multiple data sources by using a combination of Approximate Bayesian Computation (ABC) and Markov Chain Monte Carlo (MCMC). We use the method to analyse a collection of methicillin resistant Staphylococcus aureus (MRSA) isolates.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge