Pablo Romeo
BeeManc at the PLABA Track of TAC-2024: RoBERTa for task 1 and LLaMA3.1 and GPT-4o for task 2
Nov 11, 2024


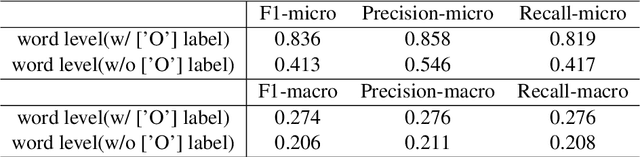
Abstract:This report is the system description of the BeeManc team for shared task Plain Language Adaptation of Biomedical Abstracts (PLABA) 2024. This report contains two sections corresponding to the two sub-tasks in PLABA 2024. In task one, we applied fine-tuned ReBERTa-Base models to identify and classify the difficult terms, jargon and acronyms in the biomedical abstracts and reported the F1 score. Due to time constraints, we didn't finish the replacement task. In task two, we leveraged Llamma3.1-70B-Instruct and GPT-4o with the one-shot prompts to complete the abstract adaptation and reported the scores in BLEU, SARI, BERTScore, LENS, and SALSA. From the official Evaluation from PLABA-2024 on Task 1A and 1B, our \textbf{much smaller fine-tuned RoBERTa-Base} model ranked 3rd and 2nd respectively on the two sub-task, and the \textbf{1st on averaged F1 scores across the two tasks} from 9 evaluated systems. Our share our fine-tuned models and related resources at \url{https://github.com/HECTA-UoM/PLABA2024}
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge