Nazmul Siddique
Ulster University, UK
BETA-Labeling for Multilingual Dataset Construction in Low-Resource IR
Feb 16, 2026Abstract:IR in low-resource languages remains limited by the scarcity of high-quality, task-specific annotated datasets. Manual annotation is expensive and difficult to scale, while using large language models (LLMs) as automated annotators introduces concerns about label reliability, bias, and evaluation validity. This work presents a Bangla IR dataset constructed using a BETA-labeling framework involving multiple LLM annotators from diverse model families. The framework incorporates contextual alignment, consistency checks, and majority agreement, followed by human evaluation to verify label quality. Beyond dataset creation, we examine whether IR datasets from other low-resource languages can be effectively reused through one-hop machine translation. Using LLM-based translation across multiple language pairs, we experimented on meaning preservation and task validity between source and translated datasets. Our experiment reveal substantial variation across languages, reflecting language-dependent biases and inconsistent semantic preservation that directly affect the reliability of cross-lingual dataset reuse. Overall, this study highlights both the potential and limitations of LLM-assisted dataset creation for low-resource IR. It provides empirical evidence of the risks associated with cross-lingual dataset reuse and offers practical guidance for constructing more reliable benchmarks and evaluation pipelines in low-resource language settings.
DL$^3$M: A Vision-to-Language Framework for Expert-Level Medical Reasoning through Deep Learning and Large Language Models
Dec 14, 2025Abstract:Medical image classifiers detect gastrointestinal diseases well, but they do not explain their decisions. Large language models can generate clinical text, yet they struggle with visual reasoning and often produce unstable or incorrect explanations. This leaves a gap between what a model sees and the type of reasoning a clinician expects. We introduce a framework that links image classification with structured clinical reasoning. A new hybrid model, MobileCoAtNet, is designed for endoscopic images and achieves high accuracy across eight stomach-related classes. Its outputs are then used to drive reasoning by several LLMs. To judge this reasoning, we build two expert-verified benchmarks covering causes, symptoms, treatment, lifestyle, and follow-up care. Thirty-two LLMs are evaluated against these gold standards. Strong classification improves the quality of their explanations, but none of the models reach human-level stability. Even the best LLMs change their reasoning when prompts vary. Our study shows that combining DL with LLMs can produce useful clinical narratives, but current LLMs remain unreliable for high-stakes medical decisions. The framework provides a clearer view of their limits and a path for building safer reasoning systems. The complete source code and datasets used in this study are available at https://github.com/souravbasakshuvo/DL3M.
Device-friendly Guava fruit and leaf disease detection using deep learning
Sep 26, 2022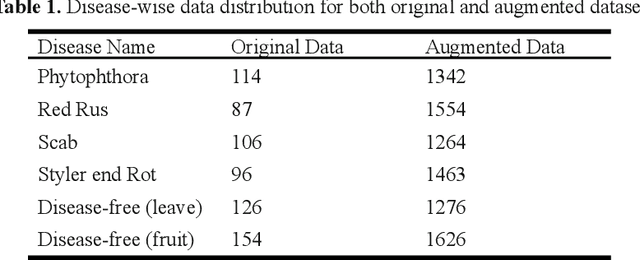
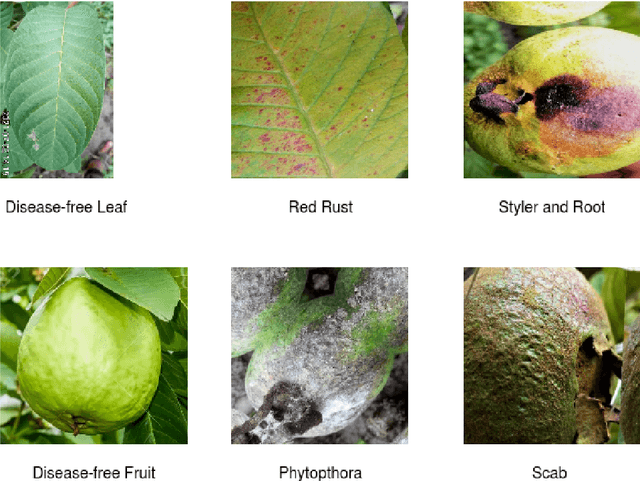

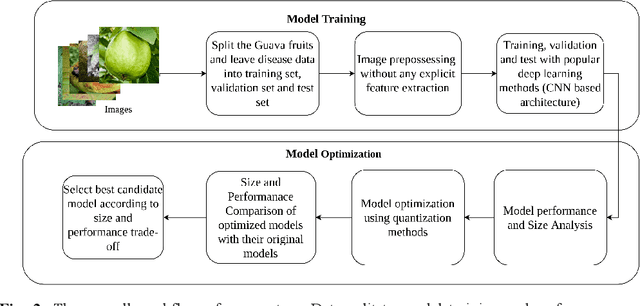
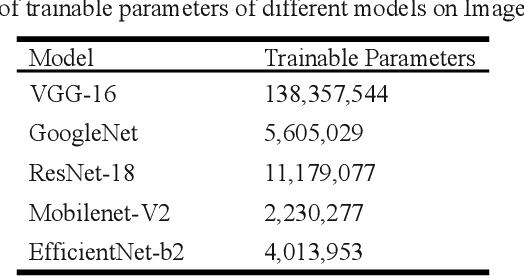
Abstract:This work presents a deep learning-based plant disease diagnostic system using images of fruits and leaves. Five state-of-the-art convolutional neural networks (CNN) have been employed for implementing the system. Hitherto model accuracy has been the focus for such applications and model optimization has not been accounted for the model to be applicable to end-user devices. Two model quantization techniques such as float16 and dynamic range quantization have been applied to the five state-of-the-art CNN architectures. The study shows that the quantized GoogleNet model achieved the size of 0.143 MB with an accuracy of 97%, which is the best candidate model considering the size criterion. The EfficientNet model achieved the size of 4.2MB with an accuracy of 99%, which is the best model considering the performance criterion. The source codes are available at https://github.com/CompostieAI/Guava-disease-detection.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge