Nataliya Portman
McConnell Brain Imaging Centre, Montreal Neurological Institute, McGill University, Montreal, QC, Canada
Local semi-supervised approach to brain tissue classification in child brain MRI
May 20, 2020
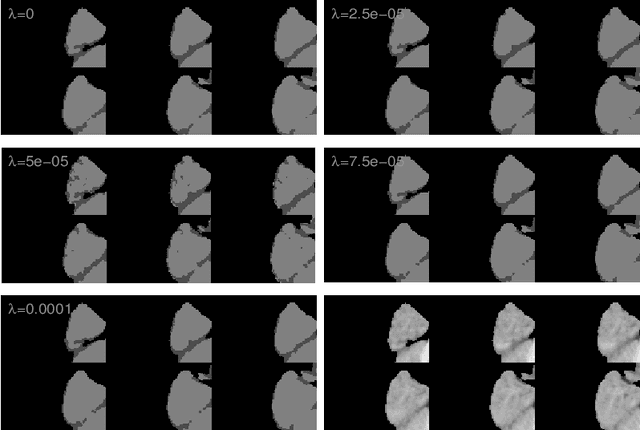

Abstract:Most segmentation methods in child brain MRI are supervised and are based on global intensity distributions of major brain structures. The successful implementation of a supervised approach depends on availability of an age-appropriate probabilistic brain atlas. For the study of early normal brain development, the construction of such a brain atlas remains a significant challenge. Moreover, using global intensity statistics leads to inaccurate detection of major brain tissue classes due to substantial intensity variations of MR signal within the constituent parts of early developing brain. In order to overcome these methodological limitations we develop a local, semi-supervised framework. It is based on Kernel Fisher Discriminant Analysis (KFDA) for pattern recognition, combined with an objective structural similarity index (SSIM) for perceptual image quality assessment. The proposed method performs optimal brain partitioning into subdomains having different average intensity values followed by SSIM-guided computation of separating surfaces between the constituent brain parts. The classified image subdomains are then stitched slice by slice via simulated annealing to form a global image of the classified brain. In this paper, we consider classification into major tissue classes (white matter and grey matter) and the cerebrospinal fluid and illustrate the proposed framework on examples of brain templates for ages 8 to 11 months and ages 44 to 60 months. We show that our method improves detection of the tissue classes by its comparison to state-of-the-art classification techniques known as Partial Volume Estimation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge