Maya Ramchandran
Cross-Cluster Weighted Forests
May 17, 2021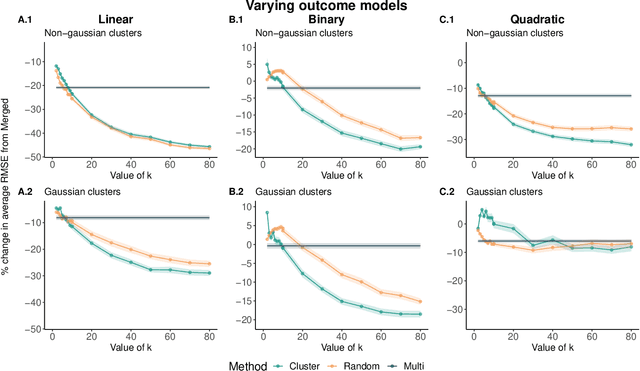
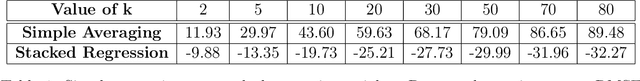
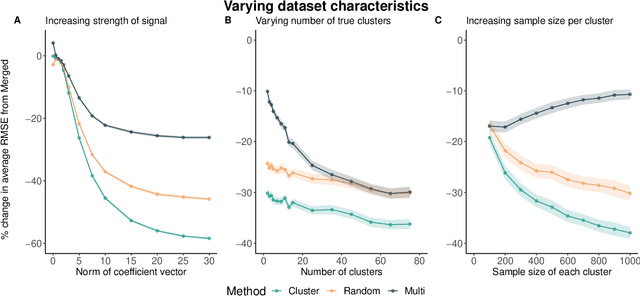
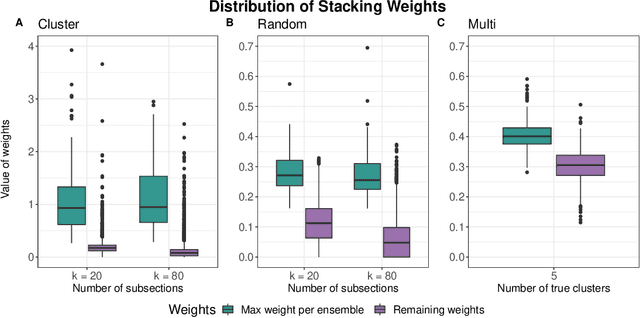
Abstract:Adapting machine learning algorithms to better handle the presence of natural clustering or batch effects within training datasets is imperative across a wide variety of biological applications. This article considers the effect of ensembling Random Forest learners trained on clusters within a single dataset with heterogeneity in the distribution of the features. We find that constructing ensembles of forests trained on clusters determined by algorithms such as k-means results in significant improvements in accuracy and generalizability over the traditional Random Forest algorithm. We denote our novel approach as the Cross-Cluster Weighted Forest, and examine its robustness to various data-generating scenarios and outcome models. Furthermore, we explore the influence of the data-partitioning and ensemble weighting strategies on conferring the benefits of our method over the existing paradigm. Finally, we apply our approach to cancer molecular profiling and gene expression datasets that are naturally divisible into clusters and illustrate that our approach outperforms classic Random Forest. Code and supplementary material are available at https://github.com/m-ramchandran/cross-cluster.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge