Maria Giulia Preti
Institute of Bioengineering, École Polytechnique Fédérale de Lausanne, Campus Biotech, Geneva, Switzerland, Department of Radiology and Medical Informatics, University of Geneva, Geneva, Switzerland
Guided Graph Spectral Embedding: Application to the C. elegans Connectome
Dec 10, 2018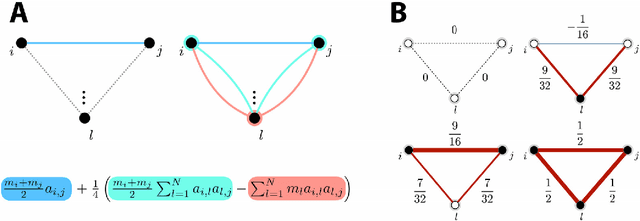
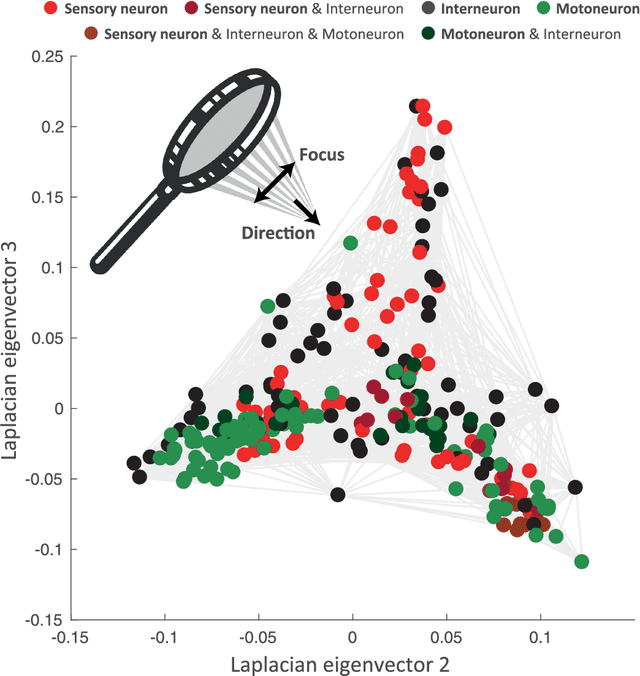
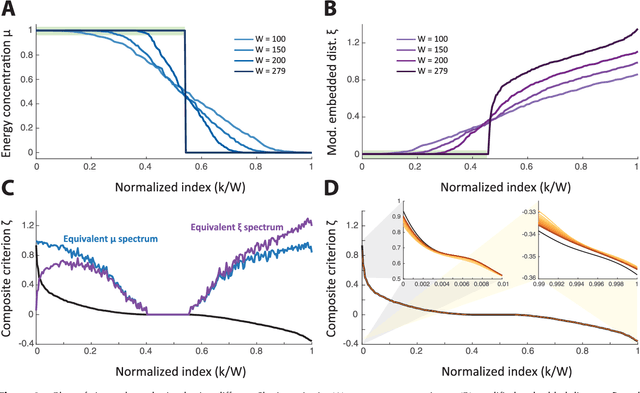
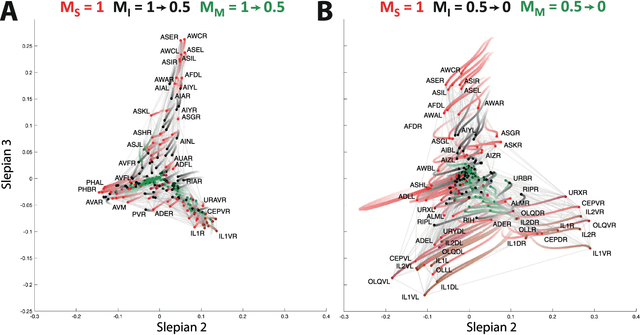
Abstract:Graph spectral analysis can yield meaningful embeddings of graphs by providing insight into distributed features not directly accessible in nodal domain. Recent efforts in graph signal processing have proposed new decompositions-e.g., based on wavelets and Slepians-that can be applied to filter signals defined on the graph. In this work, we take inspiration from these constructions to define a new guided spectral embedding that combines maximizing energy concentration with minimizing modified embedded distance for a given importance weighting of the nodes. We show these optimization goals are intrinsically opposite, leading to a well-defined and stable spectral decomposition. The importance weighting allows to put the focus on particular nodes and tune the trade-off between global and local effects. Following the derivation of our new optimization criterion and its linear approximation, we exemplify the methodology on the C. elegans structural connectome. The results of our analyses confirm known observations on the nematode's neural network in terms of functionality and importance of cells. Compared to Laplacian embedding, the guided approach, focused on a certain class of cells (sensory, inter- and motoneurons), provides more biological insights, such as the distinction between somatic positions of cells, and their involvement in low or high order processing functions.
Guiding Network Analysis using Graph Slepians: An Illustration for the C. Elegans Connectome
Aug 18, 2017Abstract:Spectral approaches of network analysis heavily rely upon the eigendecomposition of the graph Laplacian. For instance, in graph signal processing, the Laplacian eigendecomposition is used to define the graph Fourier transform and then transpose signal processing operations to graphs by implementing them in the spectral domain. Here, we build on recent work that generalized Slepian functions to the graph setting. In particular, graph Slepians are band-limited graph signals with maximal energy concentration in a given subgraph. We show how this approach can be used to guide network analysis; i.e., we propose a visualization that reveals network organization of a subgraph, but while striking a balance with global network structure. These developments are illustrated for the structural connectome of the C. Elegans.
When Slepian Meets Fiedler: Putting a Focus on the Graph Spectrum
Mar 21, 2017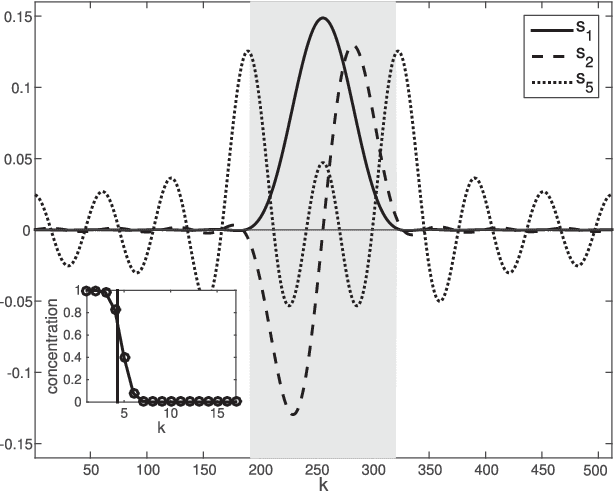
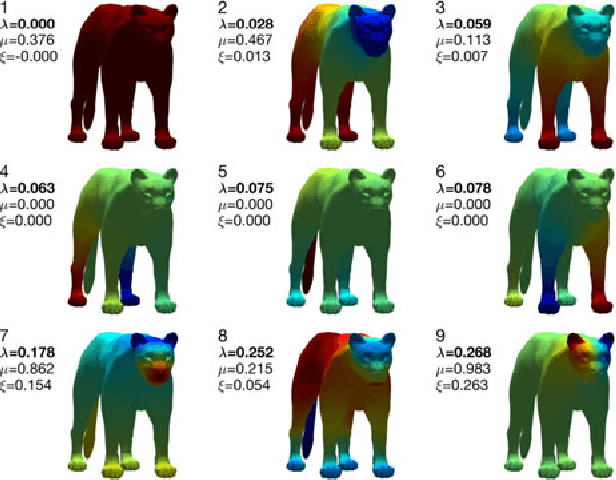
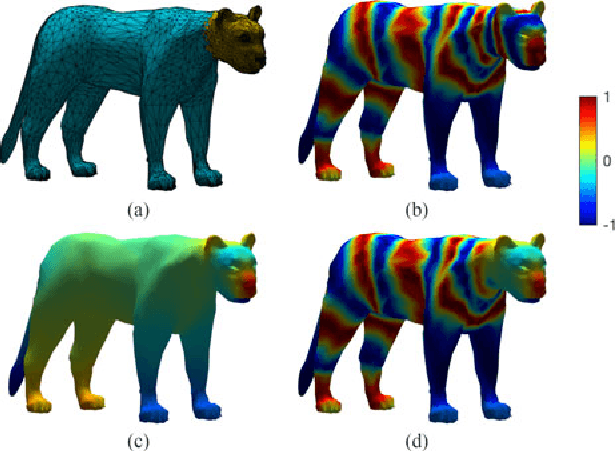


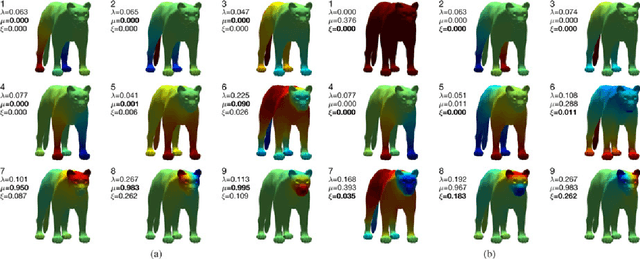
Abstract:The study of complex systems benefits from graph models and their analysis. In particular, the eigendecomposition of the graph Laplacian lets emerge properties of global organization from local interactions; e.g., the Fiedler vector has the smallest non-zero eigenvalue and plays a key role for graph clustering. Graph signal processing focusses on the analysis of signals that are attributed to the graph nodes. The eigendecomposition of the graph Laplacian allows to define the graph Fourier transform and extend conventional signal-processing operations to graphs. Here, we introduce the design of Slepian graph signals, by maximizing energy concentration in a predefined subgraph for a graph spectral bandlimit. We establish a novel link with classical Laplacian embedding and graph clustering, which provides a meaning to localized graph frequencies.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge