Manu Aggarwal
Tight basis cycle representatives for persistent homology of large data sets
Jun 06, 2022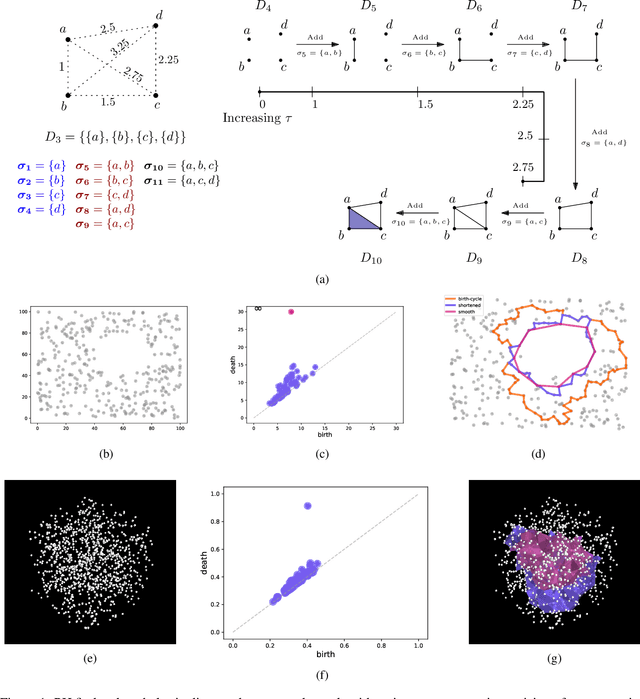

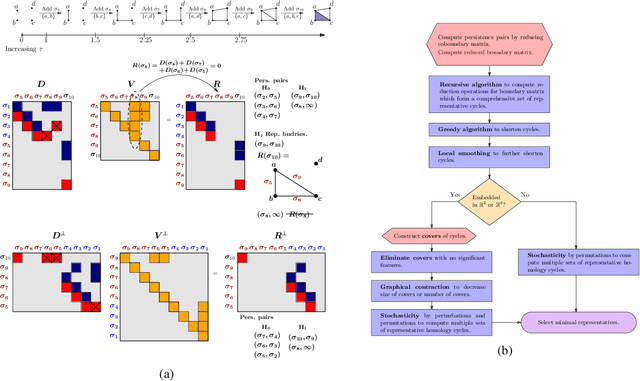
Abstract:Persistent homology (PH) is a popular tool for topological data analysis that has found applications across diverse areas of research. It provides a rigorous method to compute robust topological features in discrete experimental observations that often contain various sources of uncertainties. Although powerful in theory, PH suffers from high computation cost that precludes its application to large data sets. Additionally, most analyses using PH are limited to computing the existence of nontrivial features. Precise localization of these features is not generally attempted because, by definition, localized representations are not unique and because of even higher computation cost. For scientific applications, such a precise location is a sine qua non for determining functional significance. Here, we provide a strategy and algorithms to compute tight representative boundaries around nontrivial robust features in large data sets. To showcase the efficiency of our algorithms and the precision of computed boundaries, we analyze three data sets from different scientific fields. In the human genome, we found an unexpected effect on loops through chromosome 13 and the sex chromosomes, upon impairment of chromatin loop formation. In a distribution of galaxies in the universe, we found statistically significant voids. In protein homologs with significantly different topology, we found voids attributable to ligand-interaction, mutation, and differences between species.
Dory: Overcoming Barriers to Computing Persistent Homology
Mar 22, 2021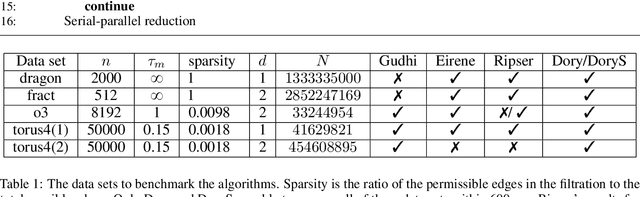
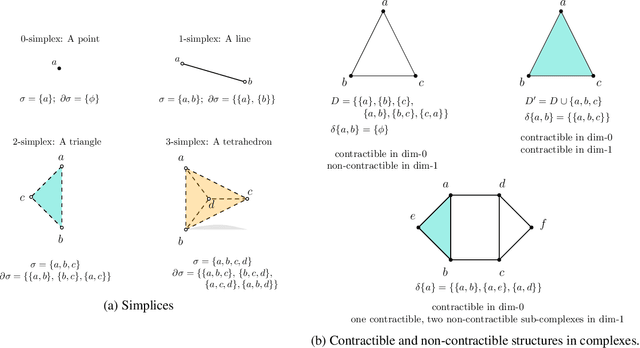
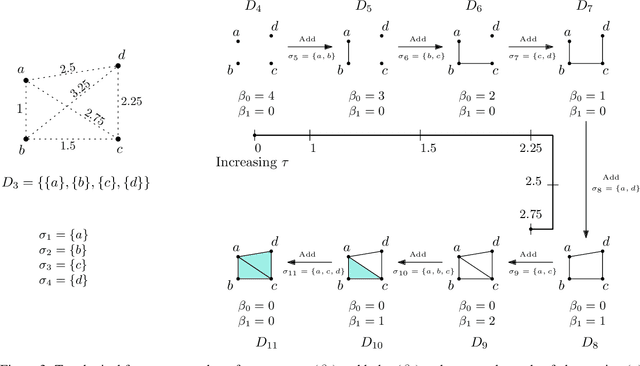
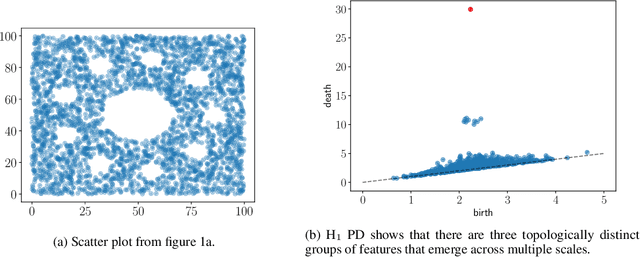
Abstract:Persistent homology (PH) is an approach to topological data analysis (TDA) that computes multi-scale topologically invariant properties of high-dimensional data that are robust to noise. While PH has revealed useful patterns across various applications, computational requirements have limited applications to small data sets of a few thousand points. We present Dory, an efficient and scalable algorithm that can compute the persistent homology of large data sets. Dory uses significantly less memory than published algorithms and also provides significant reductions in the computation time compared to most algorithms. It scales to process data sets with millions of points. As an application, we compute the PH of the human genome at high resolution as revealed by a genome-wide Hi-C data set. Results show that the topology of the human genome changes significantly upon treatment with auxin, a molecule that degrades cohesin, corroborating the hypothesis that cohesin plays a crucial role in loop formation in DNA.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge