Lucas Plagwitz
Fine-Tuning LLMs on Small Medical Datasets: Text Classification and Normalization Effectiveness on Cardiology reports and Discharge records
Mar 27, 2025Abstract:We investigate the effectiveness of fine-tuning large language models (LLMs) on small medical datasets for text classification and named entity recognition tasks. Using a German cardiology report dataset and the i2b2 Smoking Challenge dataset, we demonstrate that fine-tuning small LLMs locally on limited training data can improve performance achieving comparable results to larger models. Our experiments show that fine-tuning improves performance on both tasks, with notable gains observed with as few as 200-300 training examples. Overall, the study highlights the potential of task-specific fine-tuning of LLMs for automating clinical workflows and efficiently extracting structured data from unstructured medical text.
ExChanGeAI: An End-to-End Platform and Efficient Foundation Model for Electrocardiogram Analysis and Fine-tuning
Mar 17, 2025



Abstract:Electrocardiogram data, one of the most widely available biosignal data, has become increasingly valuable with the emergence of deep learning methods, providing novel insights into cardiovascular diseases and broader health conditions. However, heterogeneity of electrocardiogram formats, limited access to deep learning model weights and intricate algorithmic steps for effective fine-tuning for own disease target labels result in complex workflows. In this work, we introduce ExChanGeAI, a web-based end-to-end platform that streamlines the reading of different formats, pre-processing, visualization and custom machine learning with local and privacy-preserving fine-tuning. ExChanGeAI is adaptable for use on both personal computers and scalable to high performance server environments. The platform offers state-of-the-art deep learning models for training from scratch, alongside our novel open-source electrocardiogram foundation model CardX, pre-trained on over one million electrocardiograms. Evaluation across three external validation sets, including an entirely new testset extracted from routine care, demonstrate the fine-tuning capabilities of ExChanGeAI. CardX outperformed the benchmark foundation model while requiring significantly fewer parameters and lower computational resources. The platform enables users to empirically determine the most suitable model for their specific tasks based on systematic validations.The code is available at https://imigitlab.uni-muenster.de/published/exchangeai .
The Rlign Algorithm for Enhanced Electrocardiogram Analysis through R-Peak Alignment for Explainable Classification and Clustering
Jul 22, 2024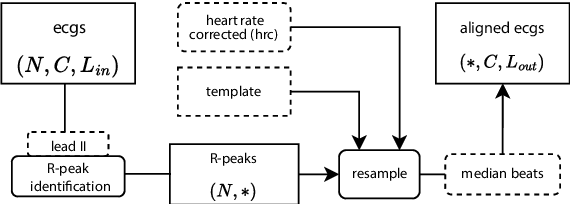
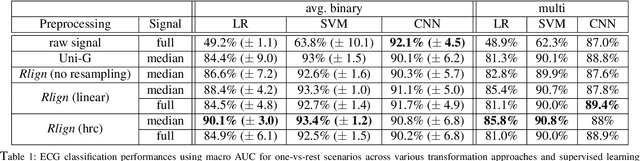
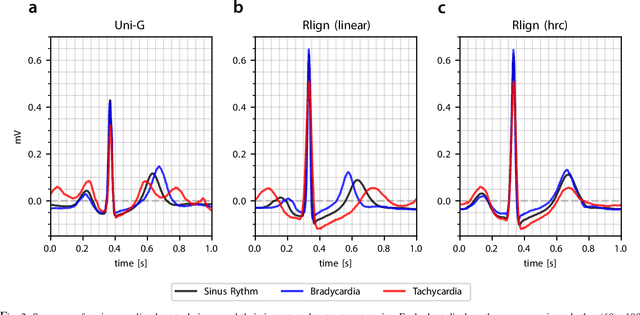
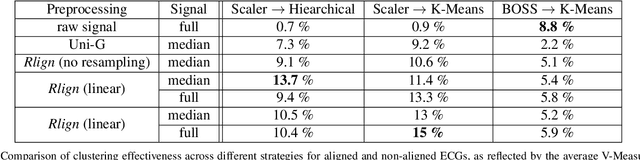
Abstract:Electrocardiogram (ECG) recordings have long been vital in diagnosing different cardiac conditions. Recently, research in the field of automatic ECG processing using machine learning methods has gained importance, mainly by utilizing deep learning methods on raw ECG signals. A major advantage of models like convolutional neural networks (CNNs) is their ability to effectively process biomedical imaging or signal data. However, this strength is tempered by challenges related to their lack of explainability, the need for a large amount of training data, and the complexities involved in adapting them for unsupervised clustering tasks. In addressing these tasks, we aim to reintroduce shallow learning techniques, including support vector machines and principal components analysis, into ECG signal processing by leveraging their semi-structured, cyclic form. To this end, we developed and evaluated a transformation that effectively restructures ECG signals into a fully structured format, facilitating their subsequent analysis using shallow learning algorithms. In this study, we present this adaptive transformative approach that aligns R-peaks across all signals in a dataset and resamples the segments between R-peaks, both with and without heart rate dependencies. We illustrate the substantial benefit of this transformation for traditional analysis techniques in the areas of classification, clustering, and explainability, outperforming commercial software for median beat transformation and CNN approaches. Our approach demonstrates a significant advantage for shallow machine learning methods over CNNs, especially when dealing with limited training data. Additionally, we release a fully tested and publicly accessible code framework, providing a robust alignment pipeline to support future research, available at https://github.com/ imi-ms/rlign.
PHOTON -- A Python API for Rapid Machine Learning Model Development
Feb 13, 2020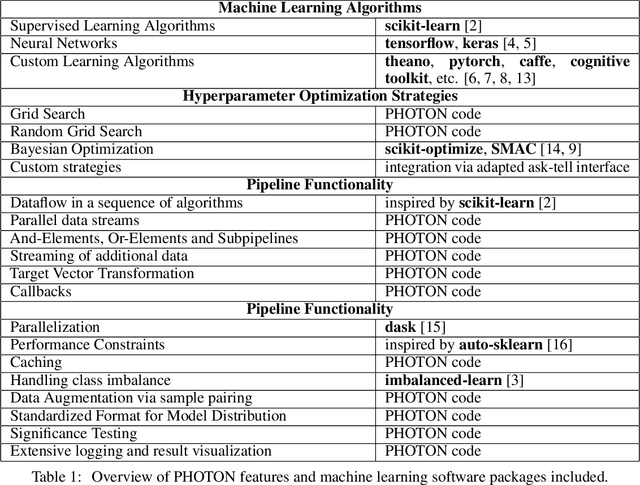
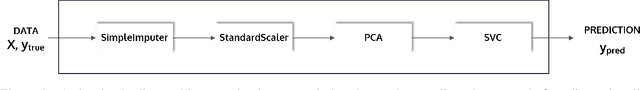
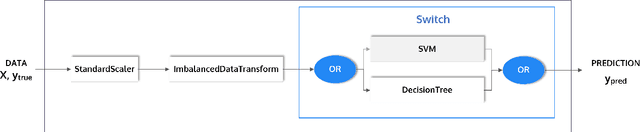
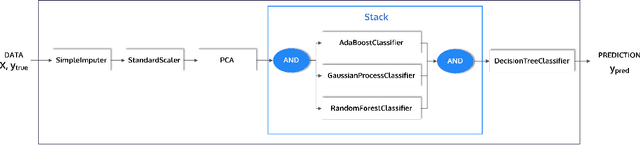
Abstract:This article describes the implementation and use of PHOTON, a high-level Python API designed to simplify and accelerate the process of machine learning model development. It enables designing both basic and advanced machine learning pipeline architectures and automatizes the repetitive training, optimization and evaluation workflow. PHOTON offers easy access to established machine learning toolboxes as well as the possibility to integrate custom algorithms and solutions for any part of the model construction and evaluation process. By adding a layer of abstraction incorporating current best practices it offers an easy-to-use, flexible approach to implementing fast, reproducible, and unbiased machine learning solutions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge