Lena Kajland Wilen
A Deep Learning Framework for Automatic Diagnosis in Lung Cancer
Jul 27, 2018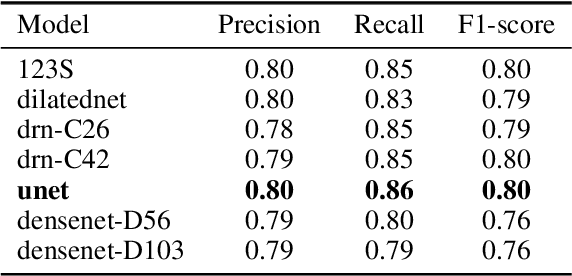

Abstract:We developed a deep learning framework that helps to automatically identify and segment lung cancer areas in patients' tissue specimens. The study was based on a cohort of lung cancer patients operated at the Uppsala University Hospital. The tissues were reviewed by lung pathologists and then the cores were compiled to tissue micro-arrays (TMAs). For experiments, hematoxylin-eosin stained slides from 712 patients were scanned and then manually annotated. Then these scans and annotations were used to train segmentation models of the developed framework. The performance of the developed deep learning framework was evaluated on fully annotated TMA cores from 178 patients reaching pixel-wise precision of 0.80 and recall of 0.86. Finally, publicly available Stanford TMA cores were used to demonstrate high performance of the framework qualitatively.
Multi-Resolution Networks for Semantic Segmentation in Whole Slide Images
Jul 25, 2018
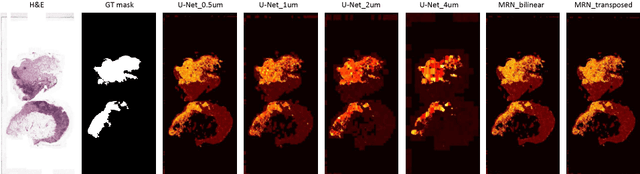



Abstract:Digital pathology provides an excellent opportunity for applying fully convolutional networks (FCNs) to tasks, such as semantic segmentation of whole slide images (WSIs). However, standard FCNs face challenges with respect to multi-resolution, inherited from the pyramid arrangement of WSIs. As a result, networks specifically designed to learn and aggregate information at different levels are desired. In this paper, we propose two novel multi-resolution networks based on the popular `U-Net' architecture, which are evaluated on a benchmark dataset for binary semantic segmentation in WSIs. The proposed methods outperform the U-Net, demonstrating superior learning and generalization capabilities.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge