Keunho Byeon
Hierarchical Classification for Improved Histopathology Image Analysis
Feb 28, 2026Abstract:Whole-slide image analysis is essential for diagnostic tasks in pathology, yet existing deep learning methods primarily rely on flat classification, ignoring hierarchical relationships among class labels. In this study, we propose HiClass, a hierarchical classification framework for improved histopathology image analysis, that enhances both coarse-grained and fine-grained WSI classification. Built based upon a multiple instance learning approach, HiClass extends it by introducing bidirectional feature integration that facilitates information exchange between coarse-grained and fine-grained feature representations, effectively learning hierarchical features. Moreover, we introduce tailored loss functions, including hierarchical consistency loss, intra- and inter-class distance loss, and group-wise cross-entropy loss, to further optimize hierarchical learning. We assess the performance of HiClass on a gastric biopsy dataset with 4 coarse-grained and 14 fine-grained classes, achieving superior classification performance for both coarse-grained classification and fine-grained classification. These results demonstrate the effectiveness of HiClass in improving WSI classification by capturing coarse-grained and fine-grained histopathological characteristics.
Pathology-Informed Latent Diffusion Model for Anomaly Detection in Lymph Node Metastasis
Aug 21, 2025Abstract:Anomaly detection is an emerging approach in digital pathology for its ability to efficiently and effectively utilize data for disease diagnosis. While supervised learning approaches deliver high accuracy, they rely on extensively annotated datasets, suffering from data scarcity in digital pathology. Unsupervised anomaly detection, however, offers a viable alternative by identifying deviations from normal tissue distributions without requiring exhaustive annotations. Recently, denoising diffusion probabilistic models have gained popularity in unsupervised anomaly detection, achieving promising performance in both natural and medical imaging datasets. Building on this, we incorporate a vision-language model with a diffusion model for unsupervised anomaly detection in digital pathology, utilizing histopathology prompts during reconstruction. Our approach employs a set of pathology-related keywords associated with normal tissues to guide the reconstruction process, facilitating the differentiation between normal and abnormal tissues. To evaluate the effectiveness of the proposed method, we conduct experiments on a gastric lymph node dataset from a local hospital and assess its generalization ability under domain shift using a public breast lymph node dataset. The experimental results highlight the potential of the proposed method for unsupervised anomaly detection across various organs in digital pathology. Code: https://github.com/QuIIL/AnoPILaD.
VLEER: Vision and Language Embeddings for Explainable Whole Slide Image Representation
Feb 28, 2025



Abstract:Recent advances in vision-language models (VLMs) have shown remarkable potential in bridging visual and textual modalities. In computational pathology, domain-specific VLMs, which are pre-trained on extensive histopathology image-text datasets, have succeeded in various downstream tasks. However, existing research has primarily focused on the pre-training process and direct applications of VLMs on the patch level, leaving their great potential for whole slide image (WSI) applications unexplored. In this study, we hypothesize that pre-trained VLMs inherently capture informative and interpretable WSI representations through quantitative feature extraction. To validate this hypothesis, we introduce Vision and Language Embeddings for Explainable WSI Representation (VLEER), a novel method designed to leverage VLMs for WSI representation. We systematically evaluate VLEER on three pathological WSI datasets, proving its better performance in WSI analysis compared to conventional vision features. More importantly, VLEER offers the unique advantage of interpretability, enabling direct human-readable insights into the results by leveraging the textual modality for detailed pathology annotations, providing clear reasoning for WSI-level pathology downstream tasks.
Benchmarking Pathology Foundation Models: Adaptation Strategies and Scenarios
Oct 21, 2024



Abstract:In computational pathology, several foundation models have recently emerged and demonstrated enhanced learning capability for analyzing pathology images. However, adapting these models to various downstream tasks remains challenging, particularly when faced with datasets from different sources and acquisition conditions, as well as limited data availability. In this study, we benchmark four pathology-specific foundation models across 14 datasets and two scenarios-consistency assessment and flexibility assessment-addressing diverse adaptation scenarios and downstream tasks. In the consistency assessment scenario, involving five fine-tuning methods, we found that the parameter-efficient fine-tuning approach was both efficient and effective for adapting pathology-specific foundation models to diverse datasets within the same downstream task. In the flexibility assessment scenario under data-limited environments, utilizing five few-shot learning methods, we observed that the foundation models benefited more from the few-shot learning methods that involve modification during the testing phase only. These findings provide insights that could guide the deployment of pathology-specific foundation models in real clinical settings, potentially improving the accuracy and reliability of pathology image analysis. The code for this study is available at: https://github.com/QuIIL/BenchmarkingPathologyFoundationModels.
CAMP: Continuous and Adaptive Learning Model in Pathology
Jul 12, 2024



Abstract:There exist numerous diagnostic tasks in pathology. Conventional computational pathology formulates and tackles them as independent and individual image classification problems, thereby resulting in computational inefficiency and high costs. To address the challenges, we propose a generic, unified, and universal framework, called a continuous and adaptive learning model in pathology (CAMP), for pathology image classification. CAMP is a generative, efficient, and adaptive classification model that can continuously adapt to any classification task by leveraging pathology-specific prior knowledge and learning taskspecific knowledge with minimal computational cost and without forgetting the knowledge from the existing tasks. We evaluated CAMP on 22 datasets, including 1,171,526 patches and 11,811 pathology slides, across 17 classification tasks. CAMP achieves state-of-theart classification performance on a wide range of datasets and tasks at both patch- and slide-levels and reduces up to 94% of computation time and 85% of storage memory in comparison to the conventional classification models. Our results demonstrate that CAMP can offer a fundamental transformation in pathology image classification, paving the way for the fully digitized and computerized pathology practice.
DIOR-ViT: Differential Ordinal Learning Vision Transformer for Cancer Classification in Pathology Images
Jul 10, 2024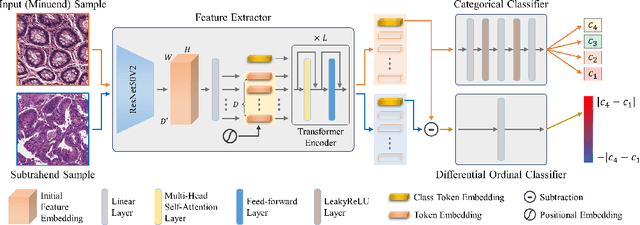



Abstract:In computational pathology, cancer grading has been mainly studied as a categorical classification problem, which does not utilize the ordering nature of cancer grades such as the higher the grade is, the worse the cancer is. To incorporate the ordering relationship among cancer grades, we introduce a differential ordinal learning problem in which we define and learn the degree of difference in the categorical class labels between pairs of samples by using their differences in the feature space. To this end, we propose a transformer-based neural network that simultaneously conducts both categorical classification and differential ordinal classification for cancer grading. We also propose a tailored loss function for differential ordinal learning. Evaluating the proposed method on three different types of cancer datasets, we demonstrate that the adoption of differential ordinal learning can improve the accuracy and reliability of cancer grading, outperforming conventional cancer grading approaches. The proposed approach should be applicable to other diseases and problems as they involve ordinal relationship among class labels.
Centroid-aware feature recalibration for cancer grading in pathology images
Jul 26, 2023



Abstract:Cancer grading is an essential task in pathology. The recent developments of artificial neural networks in computational pathology have shown that these methods hold great potential for improving the accuracy and quality of cancer diagnosis. However, the issues with the robustness and reliability of such methods have not been fully resolved yet. Herein, we propose a centroid-aware feature recalibration network that can conduct cancer grading in an accurate and robust manner. The proposed network maps an input pathology image into an embedding space and adjusts it by using centroids embedding vectors of different cancer grades via attention mechanism. Equipped with the recalibrated embedding vector, the proposed network classifiers the input pathology image into a pertinent class label, i.e., cancer grade. We evaluate the proposed network using colorectal cancer datasets that were collected under different environments. The experimental results confirm that the proposed network is able to conduct cancer grading in pathology images with high accuracy regardless of the environmental changes in the datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge