Jakub Gazda
Towards Brain MRI Foundation Models for the Clinic: Findings from the FOMO25 Challenge
Apr 13, 2026Abstract:Clinical deployment of automated brain MRI analysis faces a fundamental challenge: clinical data is heterogeneous and noisy, and high-quality labels are prohibitively costly to obtain. Self-supervised learning (SSL) can address this by leveraging the vast amounts of unlabeled data produced in clinical workflows to train robust \textit{foundation models} that adapt out-of-domain with minimal supervision. However, the development of foundation models for brain MRI has been limited by small pretraining datasets and in-domain benchmarking focused on high-quality, research-grade data. To address this gap, we organized the FOMO25 challenge as a satellite event at MICCAI 2025. FOMO25 provided participants with a large pretraining dataset, FOMO60K, and evaluated models on data sourced directly from clinical workflows in few-shot and out-of-domain settings. Tasks covered infarct classification, meningioma segmentation, and brain age regression, and considered both models trained on FOMO60K (method track) and any data (open track). Nineteen foundation models from sixteen teams were evaluated using a standardized containerized pipeline. Results show that (a) self-supervised pretraining improves generalization on clinical data under domain shift, with the strongest models trained \textit{out-of-domain} surpassing supervised baselines trained \textit{in-domain}. (b) No single pretraining objective benefits all tasks: MAE favors segmentation, hybrid reconstruction-contrastive objectives favor classification, and (c) strong performance was achieved by small pretrained models, and improvements from scaling model size and training duration did not yield reliable benefits.
Large-scale modality-invariant foundation models for brain MRI analysis: Application to lesion segmentation
Nov 14, 2025Abstract:The field of computer vision is undergoing a paradigm shift toward large-scale foundation model pre-training via self-supervised learning (SSL). Leveraging large volumes of unlabeled brain MRI data, such models can learn anatomical priors that improve few-shot performance in diverse neuroimaging tasks. However, most SSL frameworks are tailored to natural images, and their adaptation to capture multi-modal MRI information remains underexplored. This work proposes a modality-invariant representation learning setup and evaluates its effectiveness in stroke and epilepsy lesion segmentation, following large-scale pre-training. Experimental results suggest that despite successful cross-modality alignment, lesion segmentation primarily benefits from preserving fine-grained modality-specific features. Model checkpoints and code are made publicly available.
Generative Adversarial Networks in Ultrasound Imaging: Extending Field of View Beyond Conventional Limits
May 31, 2024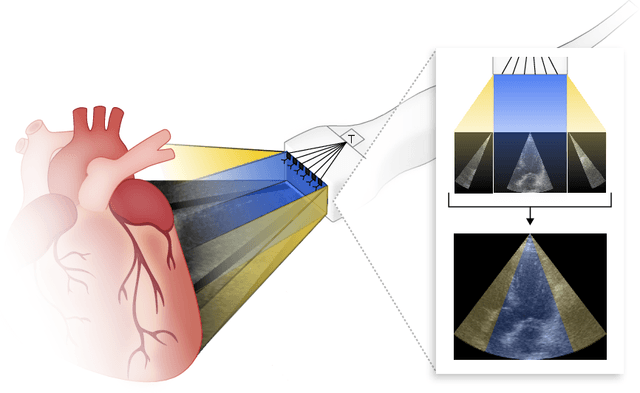
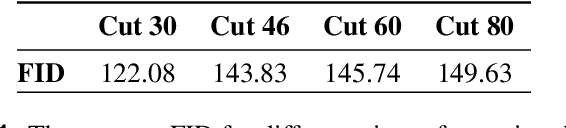
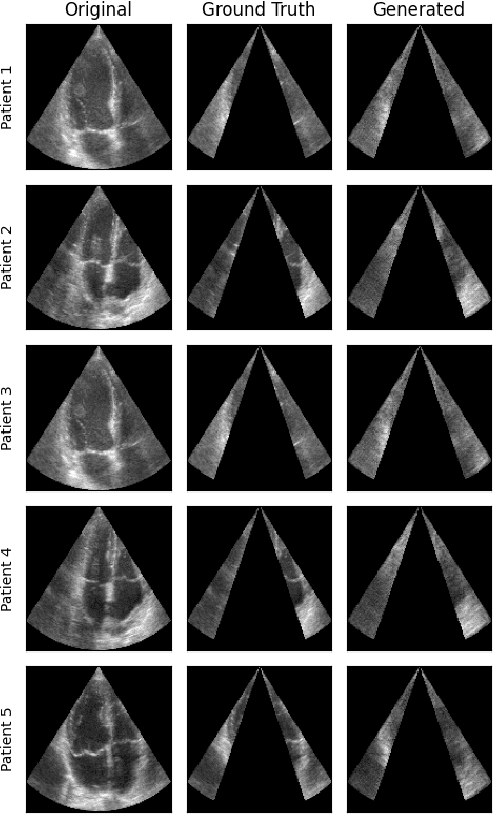
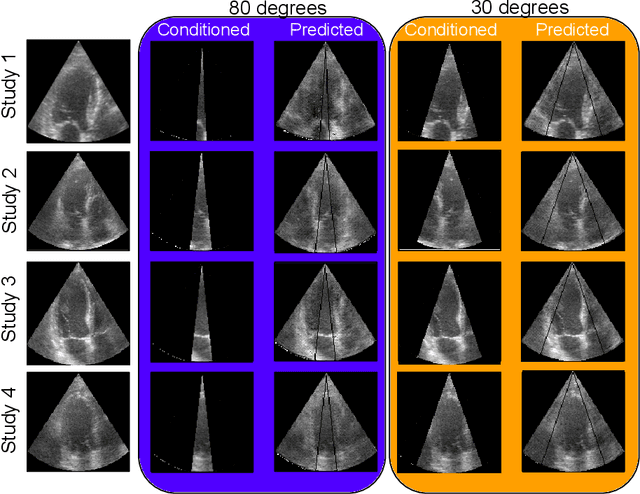
Abstract:Transthoracic Echocardiography (TTE) is a fundamental, non-invasive diagnostic tool in cardiovascular medicine, enabling detailed visualization of cardiac structures crucial for diagnosing various heart conditions. Despite its widespread use, TTE ultrasound imaging faces inherent limitations, notably the trade-off between field of view (FoV) and resolution. This paper introduces a novel application of conditional Generative Adversarial Networks (cGANs), specifically designed to extend the FoV in TTE ultrasound imaging while maintaining high resolution. Our proposed cGAN architecture, termed echoGAN, demonstrates the capability to generate realistic anatomical structures through outpainting, effectively broadening the viewable area in medical imaging. This advancement has the potential to enhance both automatic and manual ultrasound navigation, offering a more comprehensive view that could significantly reduce the learning curve associated with ultrasound imaging and aid in more accurate diagnoses. The results confirm that echoGAN reliably reproduce detailed cardiac features, thereby promising a significant step forward in the field of non-invasive cardiac naviagation and diagnostics.
Ensembled Autoencoder Regularization for Multi-Structure Segmentation for Kidney Cancer Treatment
Aug 08, 2022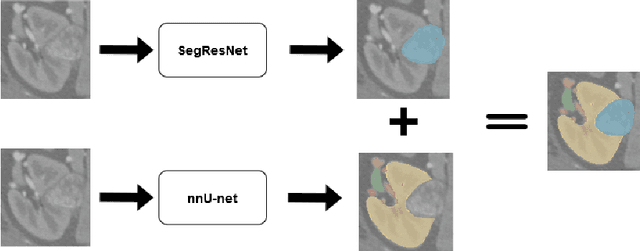
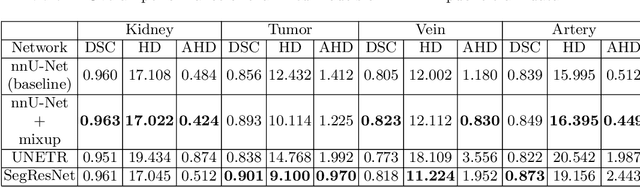
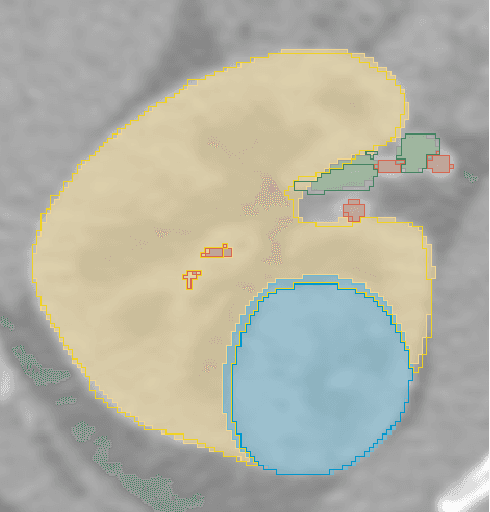

Abstract:The kidney cancer is one of the most common cancer types. The treatment frequently include surgical intervention. However, surgery is in this case particularly challenging due to regional anatomical relations. Organ delineation can significantly improve surgical planning and execution. In this contribution, we propose ensemble of two fully convolutional networks for segmentation of kidney, tumor, veins and arteries. While SegResNet architecture achieved better performance on tumor, the nnU-Net provided more precise segmentation for kidneys, arteries and veins. So in our proposed approach we combine these two networks, and further boost the performance by mixup augmentation.
Self-supervised deep convolutional neural network for chest X-ray classification
Mar 05, 2021

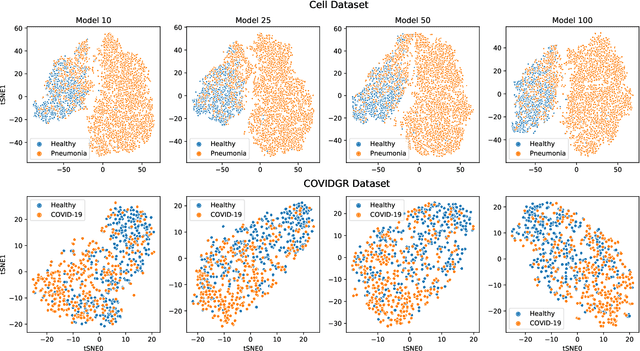
Abstract:Chest radiography is a relatively cheap, widely available medical procedure that conveys key information for making diagnostic decisions. Chest X-rays are almost always used in the diagnosis of respiratory diseases such as pneumonia or the recent COVID-19. In this paper, we propose a self-supervised deep neural network that is pretrained on an unlabeled chest X-ray dataset. The learned representations are transferred to downstream task - the classification of respiratory diseases. The results obtained on four public datasets show that our approach yields competitive results without requiring large amounts of labeled training data.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge