Heba Sailem
MesoGraph: Automatic Profiling of Malignant Mesothelioma Subtypes from Histological Images
Feb 23, 2023



Abstract:Malignant mesothelioma is classified into three histological subtypes, Epithelioid, Sarcomatoid, and Biphasic according to the relative proportions of epithelioid and sarcomatoid tumor cells present. Biphasic tumors display significant populations of both cell types. This subtyping is subjective and limited by current diagnostic guidelines and can differ even between expert thoracic pathologists when characterising the continuum of relative proportions of epithelioid and sarcomatoid components using a three class system. In this work, we develop a novel dual-task Graph Neural Network (GNN) architecture with ranking loss to learn a model capable of scoring regions of tissue down to cellular resolution. This allows quantitative profiling of a tumor sample according to the aggregate sarcomatoid association score of all the cells in the sample. The proposed approach uses only core-level labels and frames the prediction task as a dual multiple instance learning (MIL) problem. Tissue is represented by a cell graph with both cell-level morphological and regional features. We use an external multi-centric test set from Mesobank, on which we demonstrate the predictive performance of our model. We validate our model predictions through an analysis of the typical morphological features of cells according to their predicted score, finding that some of the morphological differences identified by our model match known differences used by pathologists. We further show that the model score is predictive of patient survival with a hazard ratio of 2.30. The code for the proposed approach, along with the dataset, is available at: https://github.com/measty/MesoGraph.
A Graph Based Neural Network Approach to Immune Profiling of Multiplexed Tissue Samples
Feb 01, 2022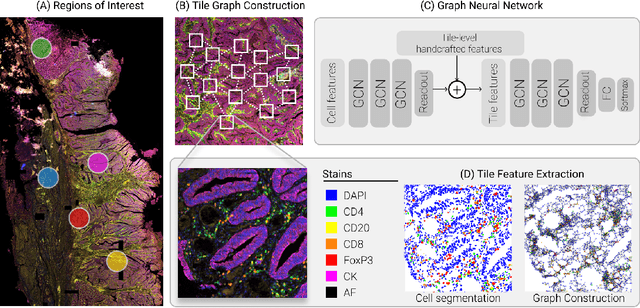
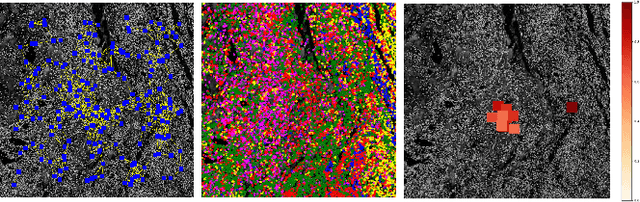
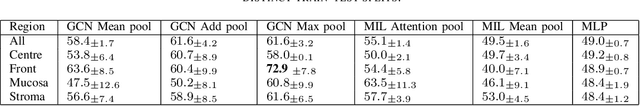
Abstract:Multiplexed immunofluorescence provides an unprecedented opportunity for studying specific cell-to-cell and cell microenvironment interactions. We employ graph neural networks to combine features obtained from tissue morphology with measurements of protein expression to profile the tumour microenvironment associated with different tumour stages. Our framework presents a new approach to analysing and processing these complex multi-dimensional datasets that overcomes some of the key challenges in analysing these data and opens up the opportunity to abstract biologically meaningful interactions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge