Geoff Tison
Conditional Synthetic Data Generation for Robust Machine Learning Applications with Limited Pandemic Data
Sep 14, 2021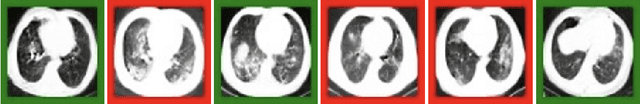

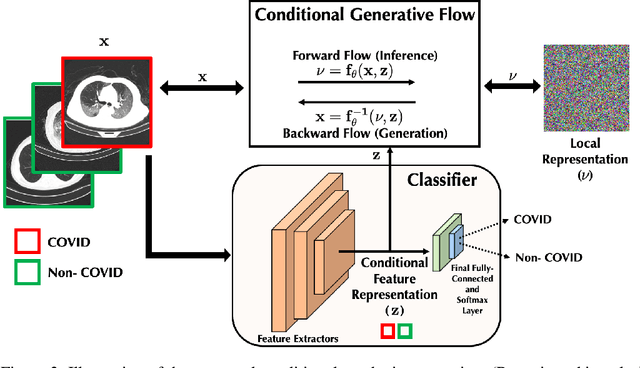
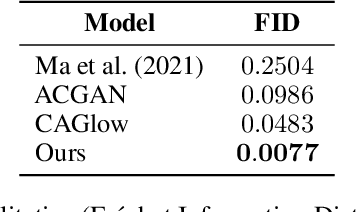
Abstract:$\textbf{Background:}$ At the onset of a pandemic, such as COVID-19, data with proper labeling/attributes corresponding to the new disease might be unavailable or sparse. Machine Learning (ML) models trained with the available data, which is limited in quantity and poor in diversity, will often be biased and inaccurate. At the same time, ML algorithms designed to fight pandemics must have good performance and be developed in a time-sensitive manner. To tackle the challenges of limited data, and label scarcity in the available data, we propose generating conditional synthetic data, to be used alongside real data for developing robust ML models. $\textbf{Methods:}$ We present a hybrid model consisting of a conditional generative flow and a classifier for conditional synthetic data generation. The classifier decouples the feature representation for the condition, which is fed to the flow to extract the local noise. We generate synthetic data by manipulating the local noise with fixed conditional feature representation. We also propose a semi-supervised approach to generate synthetic samples in the absence of labels for a majority of the available data. $\textbf{Results:}$ We performed conditional synthetic generation for chest computed tomography (CT) scans corresponding to normal, COVID-19, and pneumonia afflicted patients. We show that our method significantly outperforms existing models both on qualitative and quantitative performance, and our semi-supervised approach can efficiently synthesize conditional samples under label scarcity. As an example of downstream use of synthetic data, we show improvement in COVID-19 detection from CT scans with conditional synthetic data augmentation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge