Gaojie Li
An efficient framework based on large foundation model for cervical cytopathology whole slide image screening
Jul 16, 2024
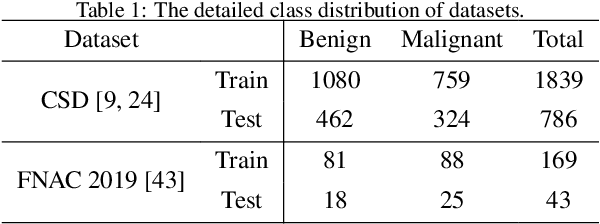
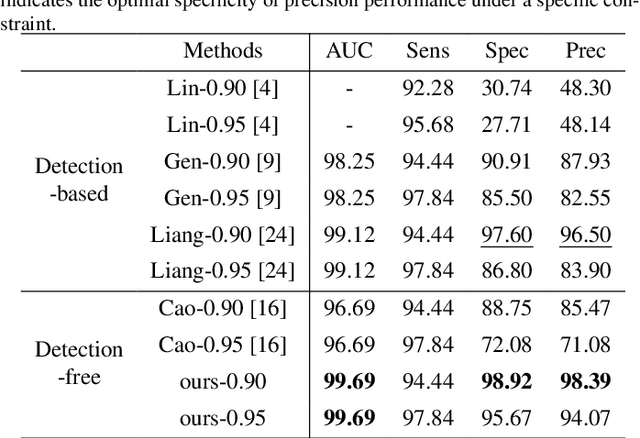
Abstract:Current cervical cytopathology whole slide image (WSI) screening primarily relies on detection-based approaches, which are limited in performance due to the expense and time-consuming annotation process. Multiple Instance Learning (MIL), a weakly supervised approach that relies solely on bag-level labels, can effectively alleviate these challenges. Nonetheless, MIL commonly employs frozen pretrained models or self-supervised learning for feature extraction, which suffers from low efficacy or inefficiency. In this paper, we propose an efficient framework for cervical cytopathology WSI classification using only WSI-level labels through unsupervised and weakly supervised learning. Given the sparse and dispersed nature of abnormal cells within cytopathological WSIs, we propose a strategy that leverages the pretrained foundation model to filter the top$k$ high-risk patches. Subsequently, we suggest parameter-efficient fine-tuning (PEFT) of a large foundation model using contrastive learning on the filtered patches to enhance its representation ability for task-specific signals. By training only the added linear adapters, we enhance the learning of patch-level features with substantially reduced time and memory consumption. Experiments conducted on the CSD and FNAC 2019 datasets demonstrate that the proposed method enhances the performance of various MIL methods and achieves state-of-the-art (SOTA) performance. The code and trained models are publicly available at https://github.com/CVIU-CSU/TCT-InfoNCE.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge