Federico Comitani
RxRx3-core: Benchmarking drug-target interactions in High-Content Microscopy
Mar 26, 2025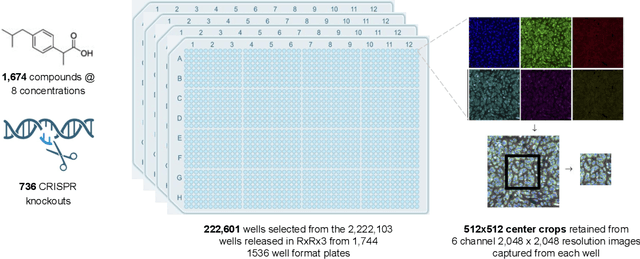
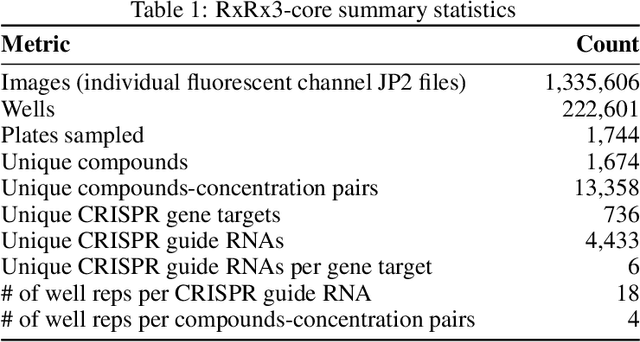


Abstract:High Content Screening (HCS) microscopy datasets have transformed the ability to profile cellular responses to genetic and chemical perturbations, enabling cell-based inference of drug-target interactions (DTI). However, the adoption of representation learning methods for HCS data has been hindered by the lack of accessible datasets and robust benchmarks. To address this gap, we present RxRx3-core, a curated and compressed subset of the RxRx3 dataset, and an associated DTI benchmarking task. At just 18GB, RxRx3-core significantly reduces the size barrier associated with large-scale HCS datasets while preserving critical data necessary for benchmarking representation learning models against a zero-shot DTI prediction task. RxRx3-core includes 222,601 microscopy images spanning 736 CRISPR knockouts and 1,674 compounds at 8 concentrations. RxRx3-core is available on HuggingFace and Polaris, along with pre-trained embeddings and benchmarking code, ensuring accessibility for the research community. By providing a compact dataset and robust benchmarks, we aim to accelerate innovation in representation learning methods for HCS data and support the discovery of novel biological insights.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge