Eamonn Kennedy
Linking Symptom Inventories using Semantic Textual Similarity
Sep 08, 2023
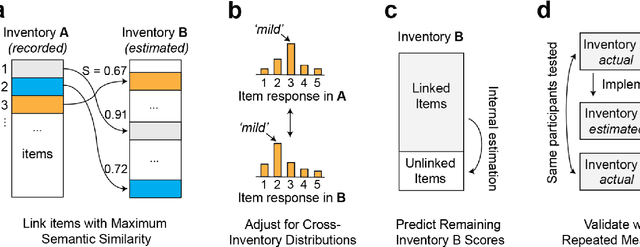
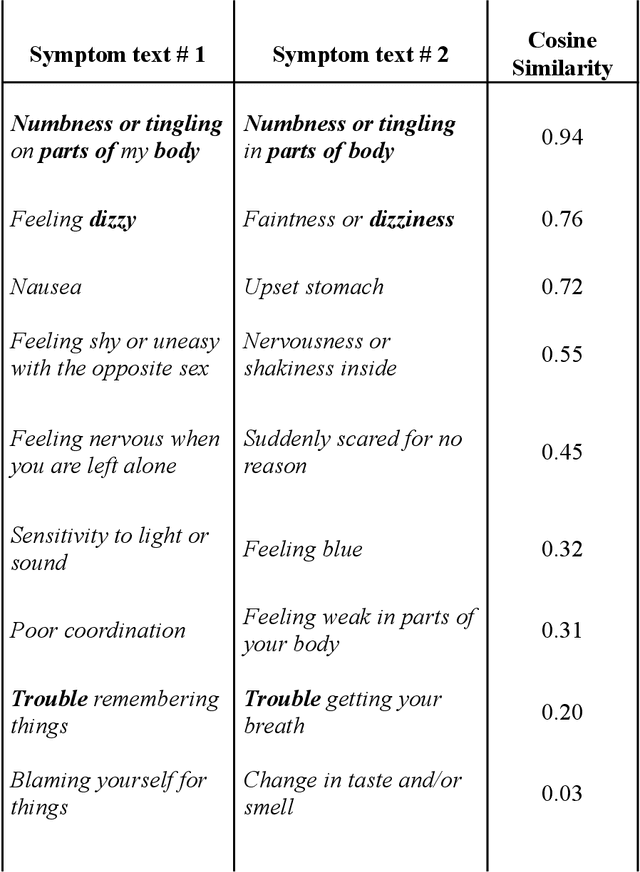
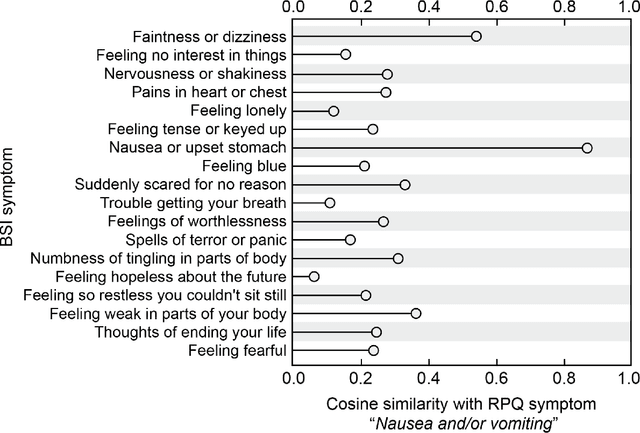
Abstract:An extensive library of symptom inventories has been developed over time to measure clinical symptoms, but this variety has led to several long standing issues. Most notably, results drawn from different settings and studies are not comparable, which limits reproducibility. Here, we present an artificial intelligence (AI) approach using semantic textual similarity (STS) to link symptoms and scores across previously incongruous symptom inventories. We tested the ability of four pre-trained STS models to screen thousands of symptom description pairs for related content - a challenging task typically requiring expert panels. Models were tasked to predict symptom severity across four different inventories for 6,607 participants drawn from 16 international data sources. The STS approach achieved 74.8% accuracy across five tasks, outperforming other models tested. This work suggests that incorporating contextual, semantic information can assist expert decision-making processes, yielding gains for both general and disease-specific clinical assessment.
Single-Molecule Protein Identification by Sub-Nanopore Sensors
Jan 09, 2017



Abstract:Recent advances in top-down mass spectrometry enabled identification of intact proteins, but this technology still faces challenges. For example, top-down mass spectrometry suffers from a lack of sensitivity since the ion counts for a single fragmentation event are often low. In contrast, nanopore technology is exquisitely sensitive to single intact molecules, but it has only been successfully applied to DNA sequencing, so far. Here, we explore the potential of sub-nanopores for single-molecule protein identification (SMPI) and describe an algorithm for identification of the electrical current blockade signal (nanospectrum) resulting from the translocation of a denaturated, linearly charged protein through a sub-nanopore. The analysis of identification p-values suggests that the current technology is already sufficient for matching nanospectra against small protein databases, e.g., protein identification in bacterial proteomes.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge