Dennis Bähr
CellCycleGAN: Spatiotemporal Microscopy Image Synthesis of Cell Populations using Statistical Shape Models and Conditional GANs
Oct 22, 2020
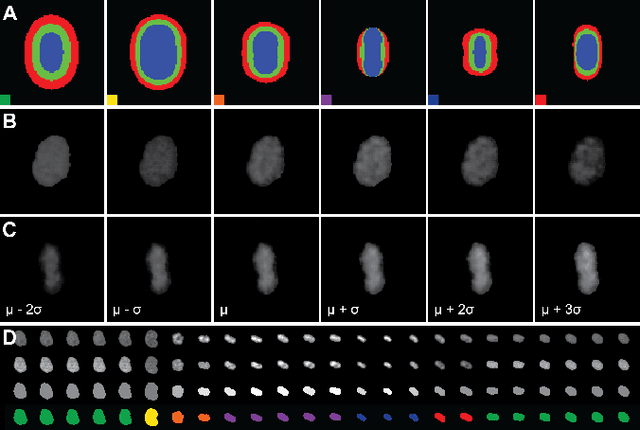
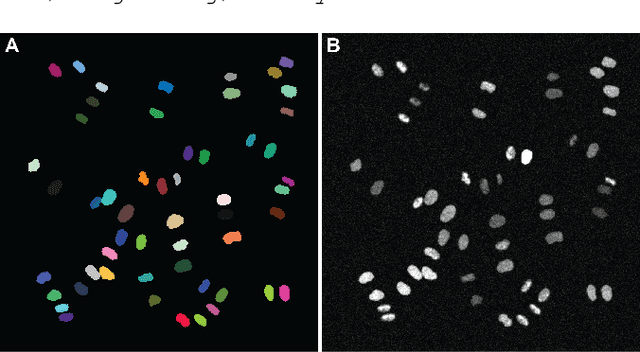
Abstract:Automatic analysis of spatio-temporal microscopy images is inevitable for state-of-the-art research in the life sciences. Recent developments in deep learning provide powerful tools for automatic analyses of such image data, but heavily depend on the amount and quality of provided training data to perform well. To this end, we developed a new method for realistic generation of synthetic 2D+t microscopy image data of fluorescently labeled cellular nuclei. The method combines spatiotemporal statistical shape models of different cell cycle stages with a conditional GAN to generate time series of cell populations and provides instance-level control of cell cycle stage and the fluorescence intensity of generated cells. We show the effect of the GAN conditioning and create a set of synthetic images that can be readily used for training and benchmarking of cell segmentation and tracking approaches.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge