Christine Decaestecker
Impact of imperfect annotations on CNN training and performance for instance segmentation and classification in digital pathology
Oct 18, 2024Abstract:Segmentation and classification of large numbers of instances, such as cell nuclei, are crucial tasks in digital pathology for accurate diagnosis. However, the availability of high-quality datasets for deep learning methods is often limited due to the complexity of the annotation process. In this work, we investigate the impact of noisy annotations on the training and performance of a state-of-the-art CNN model for the combined task of detecting, segmenting and classifying nuclei in histopathology images. In this context, we investigate the conditions for determining an appropriate number of training epochs to prevent overfitting to annotation noise during training. Our results indicate that the utilisation of a small, correctly annotated validation set is instrumental in avoiding overfitting and maintaining model performance to a large extent. Additionally, our findings underscore the beneficial role of pre-training.
Deep Learning for Reaction-Diffusion Glioma Growth Modelling: Towards a Fully Personalised Model?
Nov 26, 2021
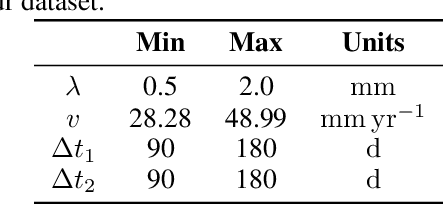


Abstract:Reaction-diffusion models have been proposed for decades to capture the growth of gliomas, the most common primary brain tumours. However, severe limitations regarding the estimation of the initial conditions and parameter values of such models have restrained their clinical use as a personalised tool. In this work, we investigate the ability of deep convolutional neural networks (DCNNs) to address the pitfalls commonly encountered in the field. Based on 1,200 synthetic tumours grown over real brain geometries derived from magnetic resonance (MR) data of 6 healthy subjects, we demonstrate the ability of DCNNs to reconstruct a whole tumour cell density distribution from only two imaging contours at a single time point. With an additional imaging contour extracted at a prior time point, we also demonstrate the ability of DCNNs to accurately estimate the individual diffusivity and proliferation parameters of the model. From this knowledge, the spatio-temporal evolution of the tumour cell density distribution at later time points can ultimately be precisely captured using the model. We finally show the applicability of our approach to MR data of a real glioblastoma patient. This approach may open the perspective of a clinical application of reaction-diffusion growth models for tumour prognosis and treatment planning.
Initial condition assessment for reaction-diffusion glioma growth models: A translational MRI/histology (in)validation study
Feb 02, 2021
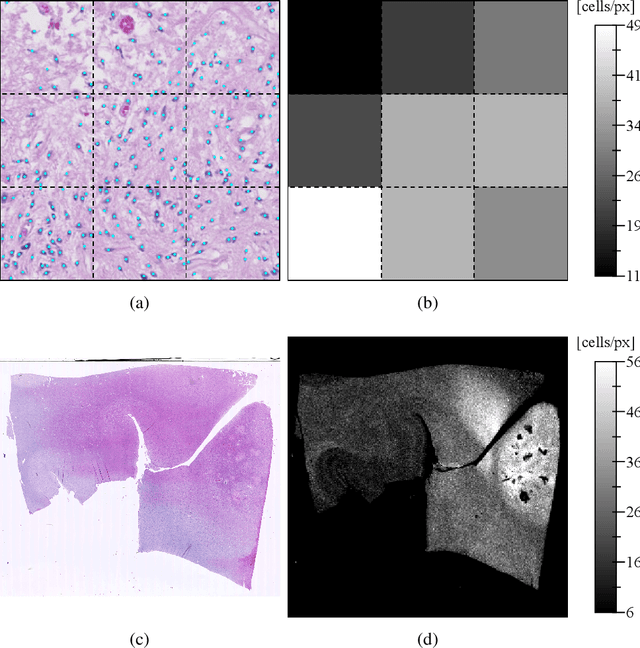

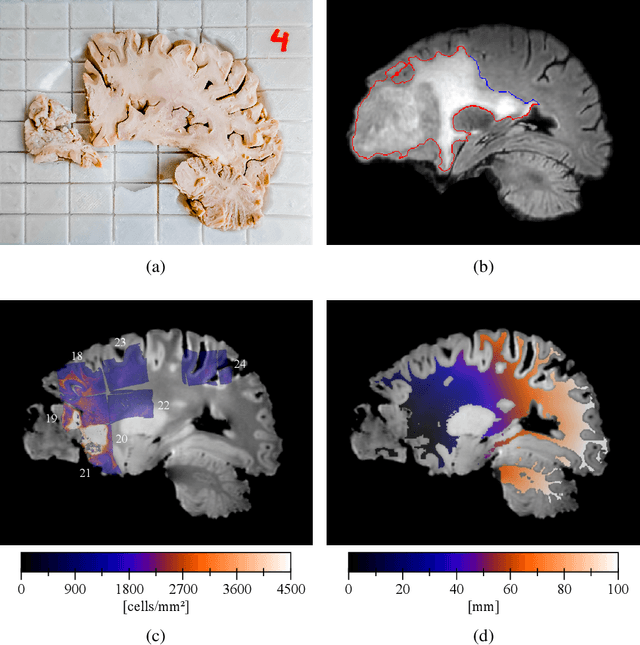
Abstract:Diffuse gliomas are highly infiltrative tumors whose early diagnosis and follow-up usually rely on magnetic resonance imaging (MRI). However, the limited sensitivity of this technique makes it impossible to directly assess the extent of the glioma cell invasion, leading to sub-optimal treatment planing. Reaction-diffusion growth models have been proposed for decades to extrapolate glioma cell infiltration beyond margins visible on MRI and predict its spatial-temporal evolution. These models nevertheless require an initial condition, that is the tumor cell density values at every location of the brain at diagnosis time. Several works have proposed to relate the tumor cell density function to abnormality outlines visible on MRI but the underlying assumptions have never been verified so far. In this work we propose to verify these assumptions by stereotactic histological analysis of a non-operated brain with glioblastoma using a tailored 3D-printed slicer. Cell density maps are computed from histological slides using a deep learning approach. The density maps are then registered to a postmortem MR image and related to an MR-derived geodesic distance map to the tumor core. The relation between the edema outlines visible on T2 FLAIR MRI and the distance to the core is also investigated. Our results suggest that (i) the previously suggested exponential decrease of the tumor cell density with the distance to the tumor core is not unreasonable but (ii) the edema outlines may in general not correspond to a cell density iso-contour and (iii) the commonly adopted tumor cell density value at these outlines is likely overestimated. These findings highlight the limitations of using conventional MRI to derive glioma cell density maps and point out the need of validating other methods to initialize reaction-diffusion growth models and make them usable in clinical practice.
Characterization of Posidonia Oceanica Seagrass Aerenchyma through Whole Slide Imaging: A Pilot Study
Mar 11, 2019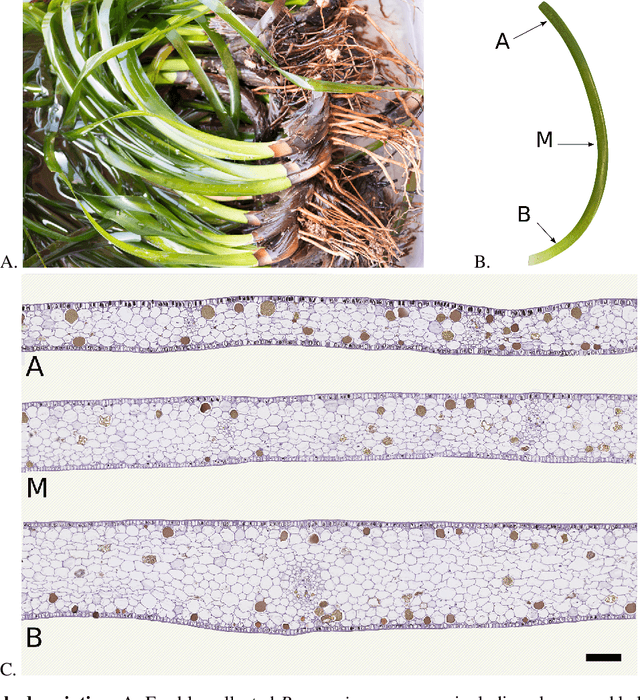
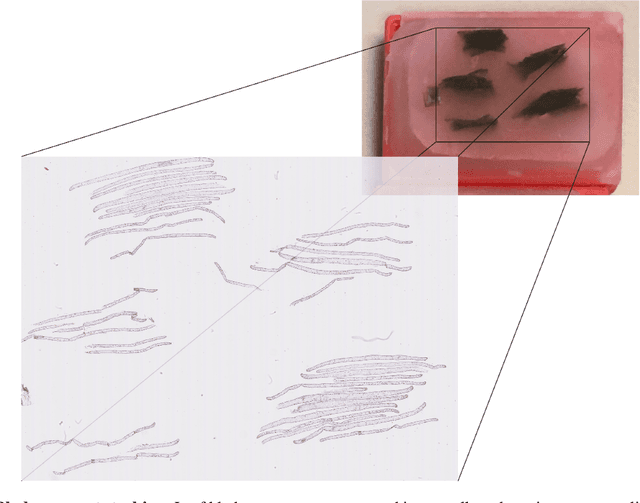
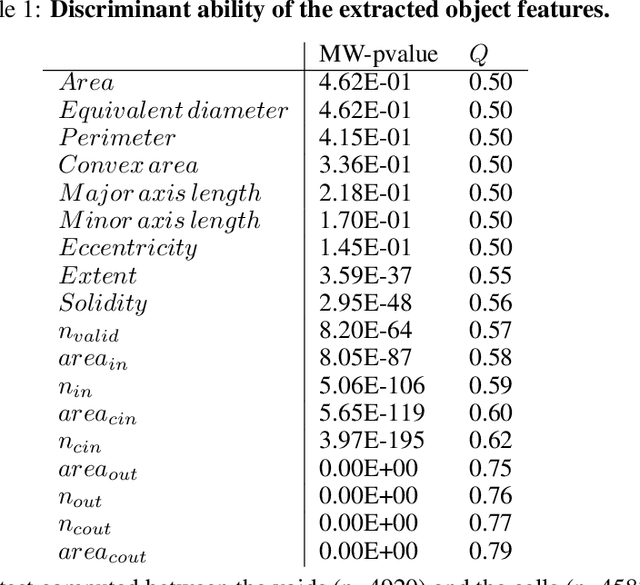


Abstract:Characterizing the tissue morphology and anatomy of seagrasses is essential to predicting their acoustic behavior. In this pilot study, we use histology techniques and whole slide imaging (WSI) to describe the composition and topology of the aerenchyma of an entire leaf blade in an automatic way combining the advantages of X-ray microtomography and optical microscopy. Paraffin blocks are prepared in such a way that microtome slices contain an arbitrarily large number of cross sections distributed along the full length of a blade. The sample organization in the paraffin block coupled with whole slide image analysis allows high throughput data extraction and an exhaustive characterization along the whole blade length. The core of the work are image processing algorithms that can identify cells and air lacunae (or void) from fiber strand, epidermis, mesophyll and vascular system. A set of specific features is developed to adequately describe the convexity of cells and voids where standard descriptors fail. The features scrutinize the local curvature of the object borders to allow an accurate discrimination between void and cell through machine learning. The algorithm allows to reconstruct the cells and cell membrane features that are relevant to tissue density, compressibility and rigidity. Size distribution of the different cell types and gas spaces, total biomass and total void volume fraction are then extracted from the high resolution slices to provide a complete characterization of the tissue along the leave from its base to the apex.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge