Catherine Helmer
BPH
Random survival forests for competing risks with multivariate longitudinal endogenous covariates
Aug 11, 2022
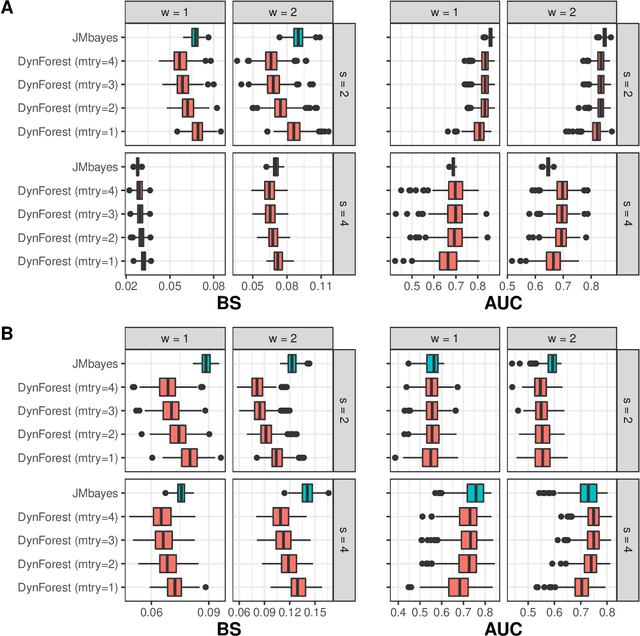
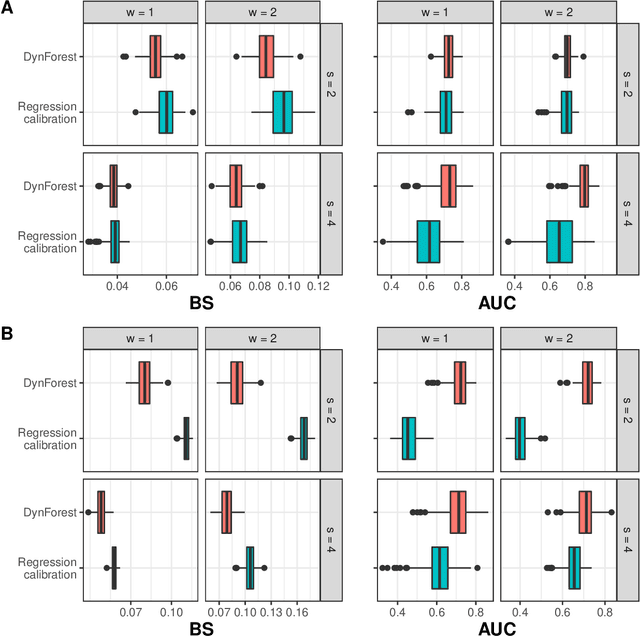
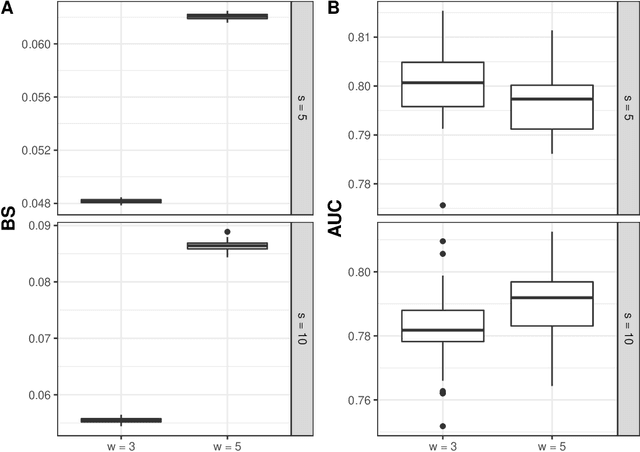
Abstract:Predicting the individual risk of a clinical event using the complete patient history is still a major challenge for personalized medicine. Among the methods developed to compute individual dynamic predictions, the joint models have the assets of using all the available information while accounting for dropout. However, they are restricted to a very small number of longitudinal predictors. Our objective was to propose an innovative alternative solution to predict an event probability using a possibly large number of longitudinal predictors. We developed DynForest, an extension of random survival forests for competing risks that handles endogenous longitudinal predictors. At each node of the trees, the time-dependent predictors are translated into time-fixed features (using mixed models) to be used as candidates for splitting the subjects into two subgroups. The individual event probability is estimated in each tree by the Aalen-Johansen estimator of the leaf in which the subject is classified according to his/her history of predictors. The final individual prediction is given by the average of the tree-specific individual event probabilities. We carried out a simulation study to demonstrate the performances of DynForest both in a small dimensional context (in comparison with joint models) and in a large dimensional context (in comparison with a regression calibration method that ignores informative dropout). We also applied DynForest to (i) predict the individual probability of dementia in the elderly according to repeated measures of cognitive, functional, vascular and neuro-degeneration markers, and (ii) quantify the importance of each type of markers for the prediction of dementia. Implemented in the R package DynForest, our methodology provides a solution for the prediction of events from longitudinal endogenous predictors whatever their number.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge