Anthony Devaux
BPH
Random Forests for time-fixed and time-dependent predictors: The DynForest R package
Feb 06, 2023
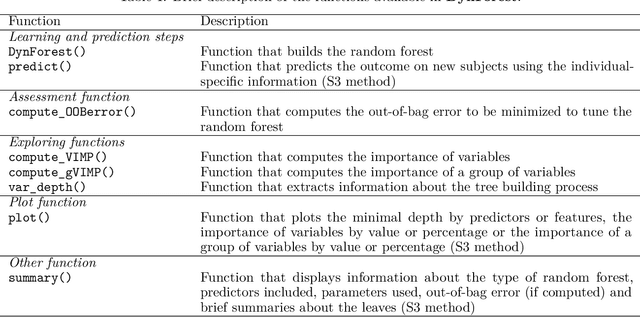



Abstract:The R package DynForest implements random forests for predicting a categorical or a (multiple causes) time-to-event outcome based on time-fixed and time-dependent predictors. Through the random forests, the time-dependent predictors can be measured with error at subject-specific times, and they can be endogeneous (i.e., impacted by the outcome process). They are modeled internally using flexible linear mixed models (thanks to lcmm package) with time-associations pre-specified by the user. DynForest computes dynamic predictions that take into account all the information from time-fixed and time-dependent predictors. DynForest also provides information about the most predictive variables using variable importance and minimal depth. Variable importance can also be computed on groups of variables. To display the results, several functions are available such as summary and plot functions. This paper aims to guide the user with a step-by-step example of the different functions for fitting random forests within DynForest.
Random survival forests for competing risks with multivariate longitudinal endogenous covariates
Aug 11, 2022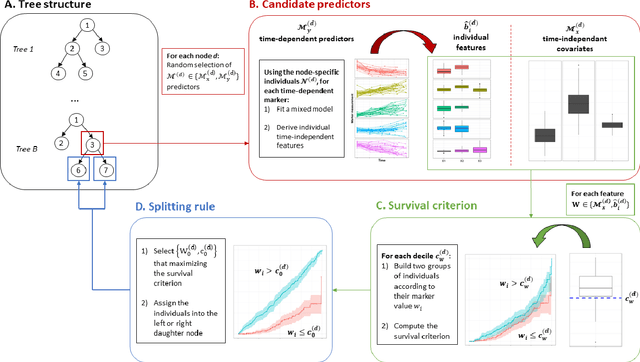
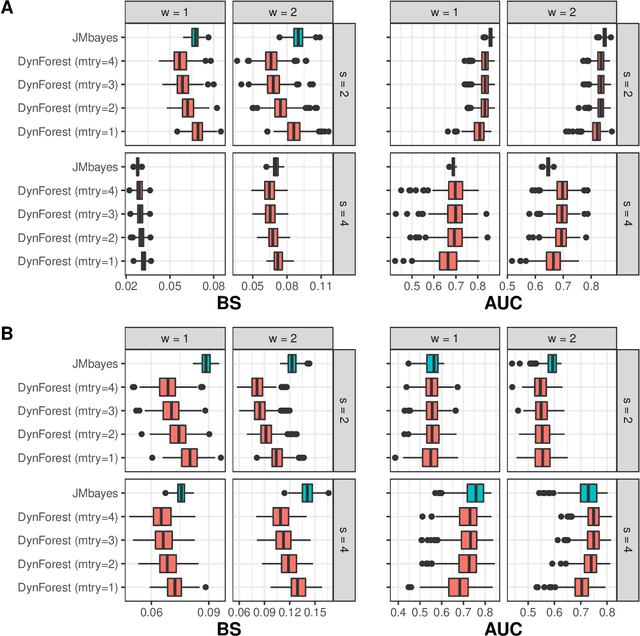
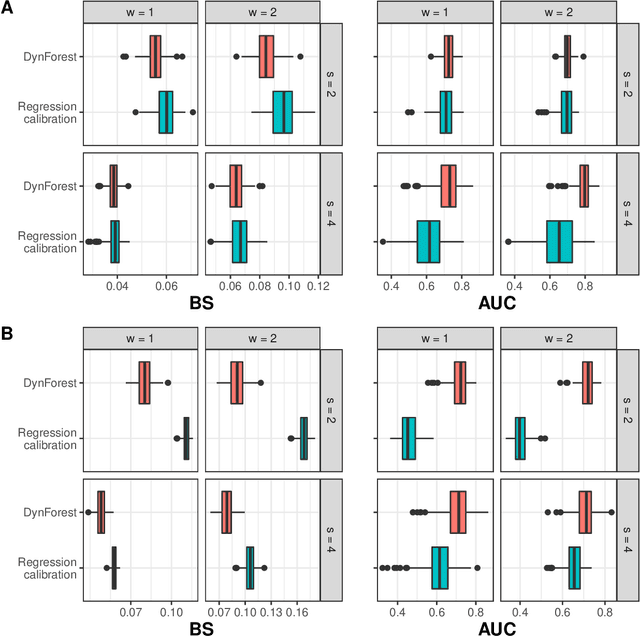
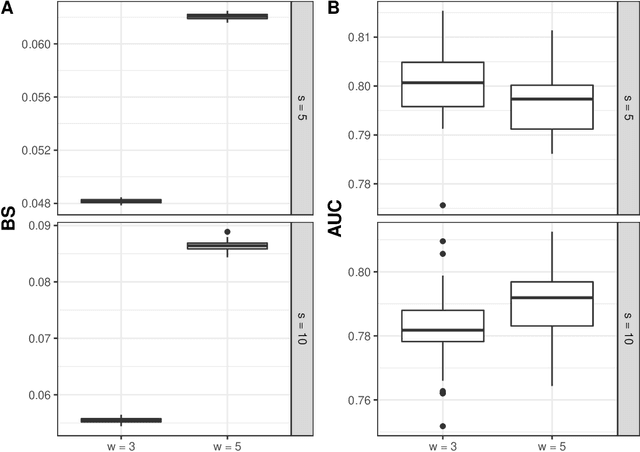
Abstract:Predicting the individual risk of a clinical event using the complete patient history is still a major challenge for personalized medicine. Among the methods developed to compute individual dynamic predictions, the joint models have the assets of using all the available information while accounting for dropout. However, they are restricted to a very small number of longitudinal predictors. Our objective was to propose an innovative alternative solution to predict an event probability using a possibly large number of longitudinal predictors. We developed DynForest, an extension of random survival forests for competing risks that handles endogenous longitudinal predictors. At each node of the trees, the time-dependent predictors are translated into time-fixed features (using mixed models) to be used as candidates for splitting the subjects into two subgroups. The individual event probability is estimated in each tree by the Aalen-Johansen estimator of the leaf in which the subject is classified according to his/her history of predictors. The final individual prediction is given by the average of the tree-specific individual event probabilities. We carried out a simulation study to demonstrate the performances of DynForest both in a small dimensional context (in comparison with joint models) and in a large dimensional context (in comparison with a regression calibration method that ignores informative dropout). We also applied DynForest to (i) predict the individual probability of dementia in the elderly according to repeated measures of cognitive, functional, vascular and neuro-degeneration markers, and (ii) quantify the importance of each type of markers for the prediction of dementia. Implemented in the R package DynForest, our methodology provides a solution for the prediction of events from longitudinal endogenous predictors whatever their number.
Individual dynamic prediction of clinical endpoint from large dimensional longitudinal biomarker history: a landmark approach
Feb 02, 2021
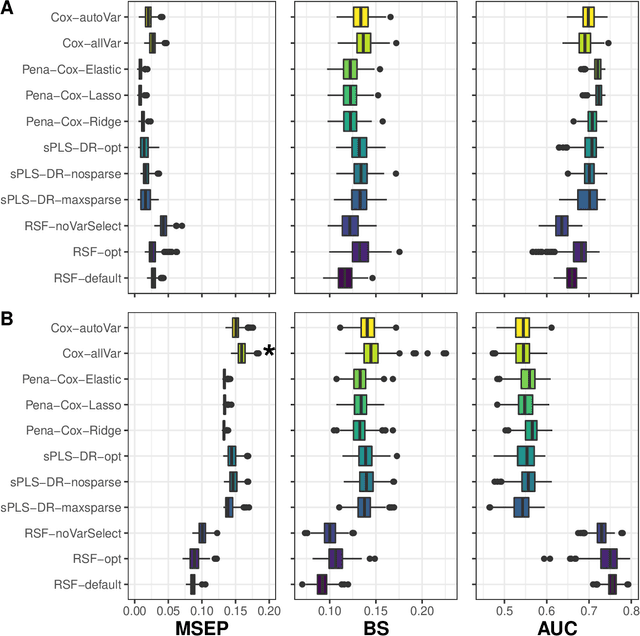
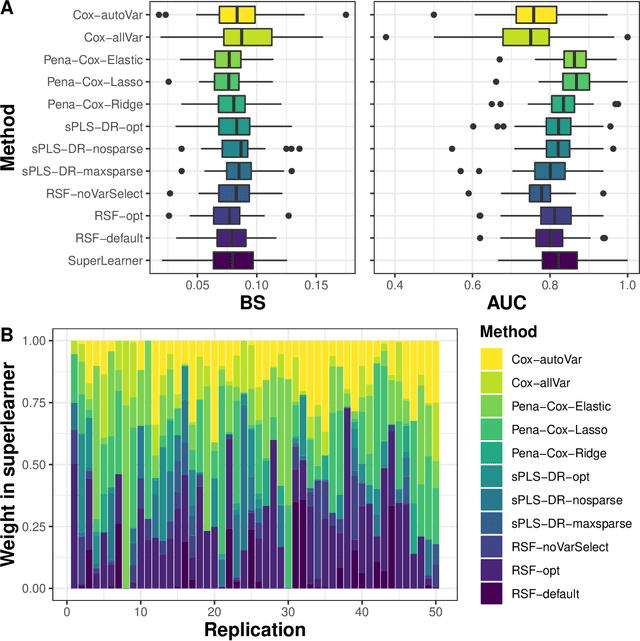
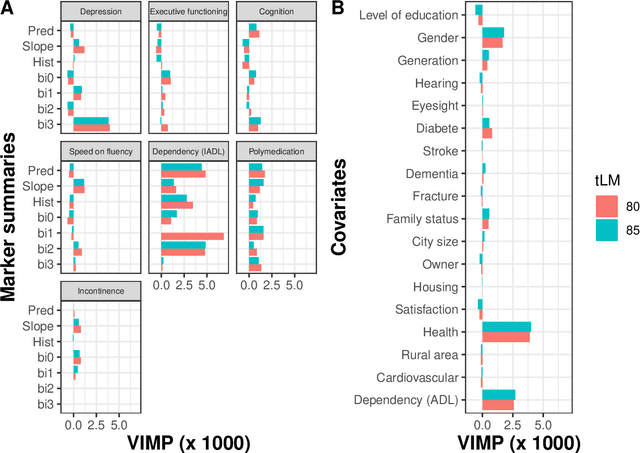
Abstract:The individual data collected throughout patient follow-up constitute crucial information for assessing the risk of a clinical event, and eventually for adapting a therapeutic strategy. Joint models and landmark models have been proposed to compute individual dynamic predictions from repeated measures to one or two markers. However, they hardly extend to the case where the complete patient history includes much more repeated markers possibly. Our objective was thus to propose a solution for the dynamic prediction of a health event that may exploit repeated measures of a possibly large number of markers. We combined a landmark approach extended to endogenous markers history with machine learning methods adapted to survival data. Each marker trajectory is modeled using the information collected up to landmark time, and summary variables that best capture the individual trajectories are derived. These summaries and additional covariates are then included in different prediction methods. To handle a possibly large dimensional history, we rely on machine learning methods adapted to survival data, namely regularized regressions and random survival forests, to predict the event from the landmark time, and we show how they can be combined into a superlearner. Then, the performances are evaluated by cross-validation using estimators of Brier Score and the area under the Receiver Operating Characteristic curve adapted to censored data. We demonstrate in a simulation study the benefits of machine learning survival methods over standard survival models, especially in the case of numerous and/or nonlinear relationships between the predictors and the event. We then applied the methodology in two prediction contexts: a clinical context with the prediction of death for patients with primary biliary cholangitis, and a public health context with the prediction of death in the general elderly population at different ages. Our methodology, implemented in R, enables the prediction of an event using the entire longitudinal patient history, even when the number of repeated markers is large. Although introduced with mixed models for the repeated markers and methods for a single right censored time-to-event, our method can be used with any other appropriate modeling technique for the markers and can be easily extended to competing risks setting.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge