BahaaEddin Alaila
Lightweight and Scalable Particle Tracking and Motion Clustering of 3D Cell Trajectories
Aug 14, 2019
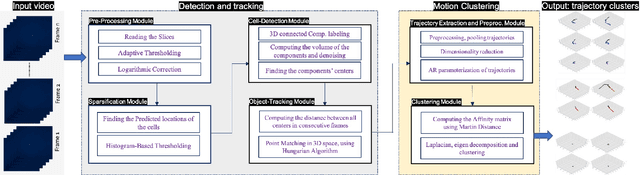
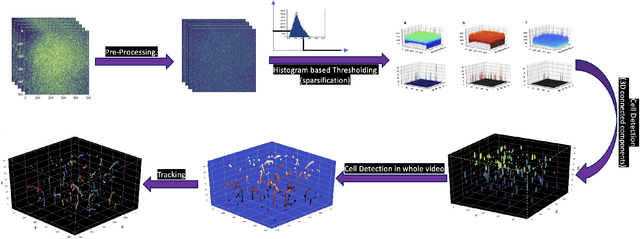
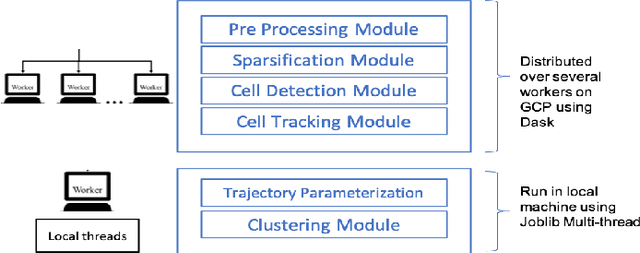
Abstract:Tracking cell particles in 3D microscopy videos is a challenging task but is of great significance for modeling the motion of cells. Proper characterization of the cell's shape, evolution, and their movement over time is crucial to understanding and modeling the mechanobiology of cell migration in many diseases. One in particular, toxoplasmosis is the disease caused by the parasite Toxoplasma gondii. Roughly, one-third of the world's population tests positive for T. gondii. Its virulence is linked to its lytic cycle, predicated on its motility and ability to enter and exit nucleated cells; therefore, studies elucidating its motility patterns are critical to the eventual development of therapeutic strategies. Here, we present a computational framework for fast and scalable detection, tracking, and identification of T. gondii motion phenotypes in 3D videos, in a completely unsupervised fashion. Our pipeline consists of several different modules including preprocessing, sparsification, cell detection, cell tracking, trajectories extraction, parametrization of the trajectories; and finally, a clustering step. Additionally, we identified the computational bottlenecks, and developed a lightweight and highly scalable pipeline through a combination of task distribution and parallelism. Our results prove both the accuracy and performance of our method.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge