Arjun Puranik
HypUC: Hyperfine Uncertainty Calibration with Gradient-boosted Corrections for Reliable Regression on Imbalanced Electrocardiograms
Nov 23, 2023Abstract:The automated analysis of medical time series, such as the electrocardiogram (ECG), electroencephalogram (EEG), pulse oximetry, etc, has the potential to serve as a valuable tool for diagnostic decisions, allowing for remote monitoring of patients and more efficient use of expensive and time-consuming medical procedures. Deep neural networks (DNNs) have been demonstrated to process such signals effectively. However, previous research has primarily focused on classifying medical time series rather than attempting to regress the continuous-valued physiological parameters central to diagnosis. One significant challenge in this regard is the imbalanced nature of the dataset, as a low prevalence of abnormal conditions can lead to heavily skewed data that results in inaccurate predictions and a lack of certainty in such predictions when deployed. To address these challenges, we propose HypUC, a framework for imbalanced probabilistic regression in medical time series, making several contributions. (i) We introduce a simple kernel density-based technique to tackle the imbalanced regression problem with medical time series. (ii) Moreover, we employ a probabilistic regression framework that allows uncertainty estimation for the predicted continuous values. (iii) We also present a new approach to calibrate the predicted uncertainty further. (iv) Finally, we demonstrate a technique to use calibrated uncertainty estimates to improve the predicted continuous value and show the efficacy of the calibrated uncertainty estimates to flag unreliable predictions. HypUC is evaluated on a large, diverse, real-world dataset of ECGs collected from millions of patients, outperforming several conventional baselines on various diagnostic tasks, suggesting a potential use-case for the reliable clinical deployment of deep learning models.
* Published at TMLR
Augmented Curation of Unstructured Clinical Notes from a Massive EHR System Reveals Specific Phenotypic Signature of Impending COVID-19 Diagnosis
Apr 28, 2020
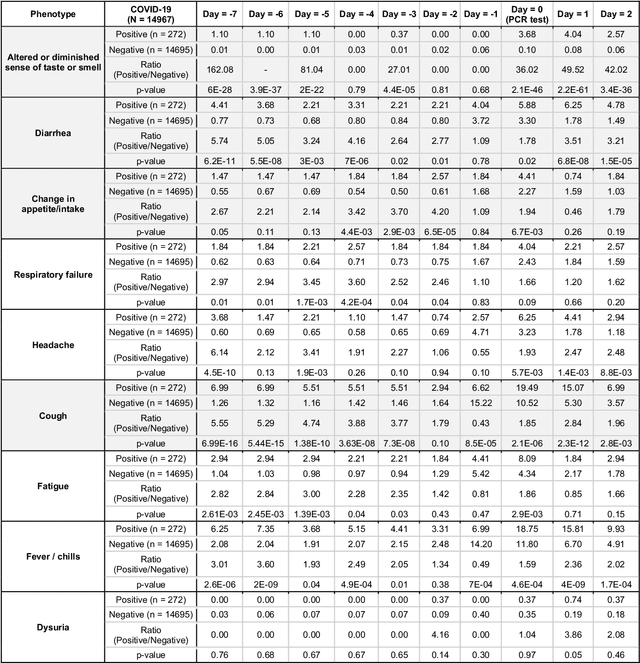
Abstract:Understanding the temporal dynamics of COVID-19 patient phenotypes is necessary to derive fine-grained resolution of pathophysiology. Here we use state-of-the-art deep neural networks over an institution-wide machine intelligence platform for the augmented curation of 15.8 million clinical notes from 30,494 patients subjected to COVID-19 PCR diagnostic testing. By contrasting the Electronic Health Record (EHR)-derived clinical phenotypes of COVID-19-positive (COVIDpos, n=635) versus COVID-19-negative (COVIDneg, n=29,859) patients over each day of the week preceding the PCR testing date, we identify anosmia/dysgeusia (37.4-fold), myalgia/arthralgia (2.6-fold), diarrhea (2.2-fold), fever/chills (2.1-fold), respiratory difficulty (1.9-fold), and cough (1.8-fold) as significantly amplified in COVIDpos over COVIDneg patients. The specific combination of cough and diarrhea has a 3.2-fold amplification in COVIDpos patients during the week prior to PCR testing, and along with anosmia/dysgeusia, constitutes the earliest EHR-derived signature of COVID-19 (4-7 days prior to typical PCR testing date). This study introduces an Augmented Intelligence platform for the real-time synthesis of institutional knowledge captured in EHRs. The platform holds tremendous potential for scaling up curation throughput, with minimal need for retraining underlying neural networks, thus promising EHR-powered early diagnosis for a broad spectrum of diseases.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge