Andrey Solovyev
DRIVE: Machine Learning to Identify Drivers of Cancer with High-Dimensional Genomic Data & Imputed Labels
May 02, 2021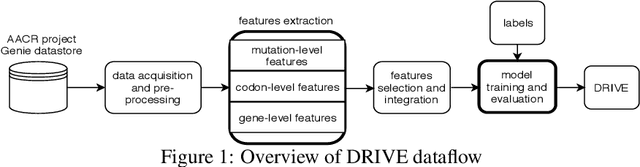
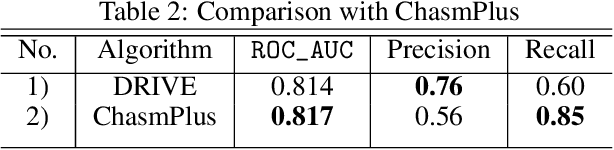
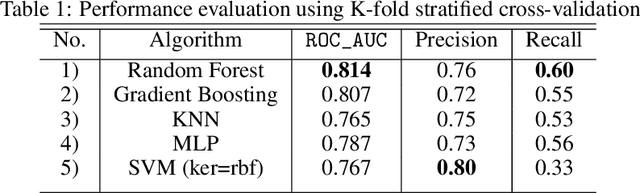
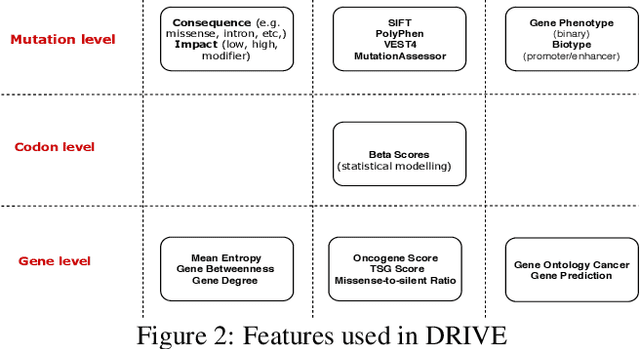

Abstract:Identifying the mutations that drive cancer growth is key in clinical decision making and precision oncology. As driver mutations confer selective advantage and thus have an increased likelihood of occurrence, frequency-based statistical models are currently favoured. These methods are not suited to rare, low frequency, driver mutations. The alternative approach to address this is through functional-impact scores, however methods using this approach are highly prone to false positives. In this paper, we propose a novel combination method for driver mutation identification, which uses the power of both statistical modelling and functional-impact based methods. Initial results show this approach outperforms the state-of-the-art methods in terms of precision, and provides comparable performance in terms of area under receiver operating characteristic curves (AU-ROC). We believe that data-driven systems based on machine learning, such as these, will become an integral part of precision oncology in the near future.
Effective Sub-clonal Cancer Representation to Predict Tumor Evolution
Nov 28, 2019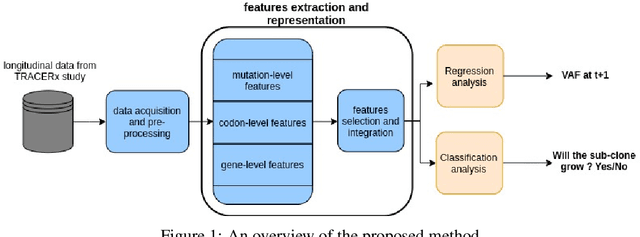
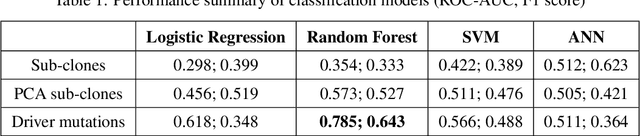
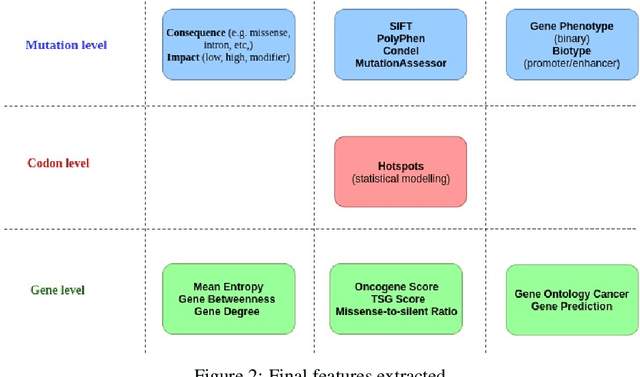
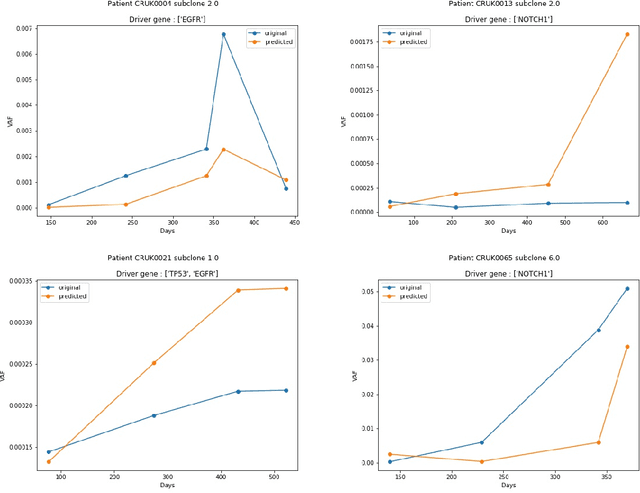
Abstract:The majority of cancer treatments end in failure due to Intra-Tumor Heterogeneity (ITH). ITH in cancer is represented by clonal evolution where different sub-clones compete with each other for resources under conditions of Darwinian natural selection. Predicting the growth of these sub-clones within a tumour is among the key challenges of modern cancer research. Predicting tumor behavior enables the creation of risk profiles for patients and the optimisation of their treatment by therapeutically targeting sub-clones more likely to grow. Current research efforts in this space are focused on mathematical modelling of population genetics to quantify the selective advantage of sub-clones, thus enabling predictions of which sub-clones are more likely to grow. These tumor evolution models are based on assumptions which are not valid for real-world tumor micro-environment. Furthermore, these models are often fit on a single instance of a tumor, and hence prediction models cannot be validated. This paper presents an alternative approach for predicting cancer evolution using a data-driven machine learning method. Our proposed method is based on the intuition that if we can capture the true characteristics of sub-clones within a tumor and represent it in the form of features, a sophisticated machine learning algorithm can be trained to predict its behavior. The work presented here provides a novel approach to predicting cancer evolution, utilizing a data-driver approach. We strongly believe that the accumulation of data from microbiologists, oncologists and machine learning researchers could be used to encapsulate the true essence of tumor sub-clones, and can play a vital role in selecting the best cancer treatments for patients.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge