Anant Shinde
Meta-Analysis of Transfer Learning for Segmentation of Brain Lesions
Jun 20, 2023
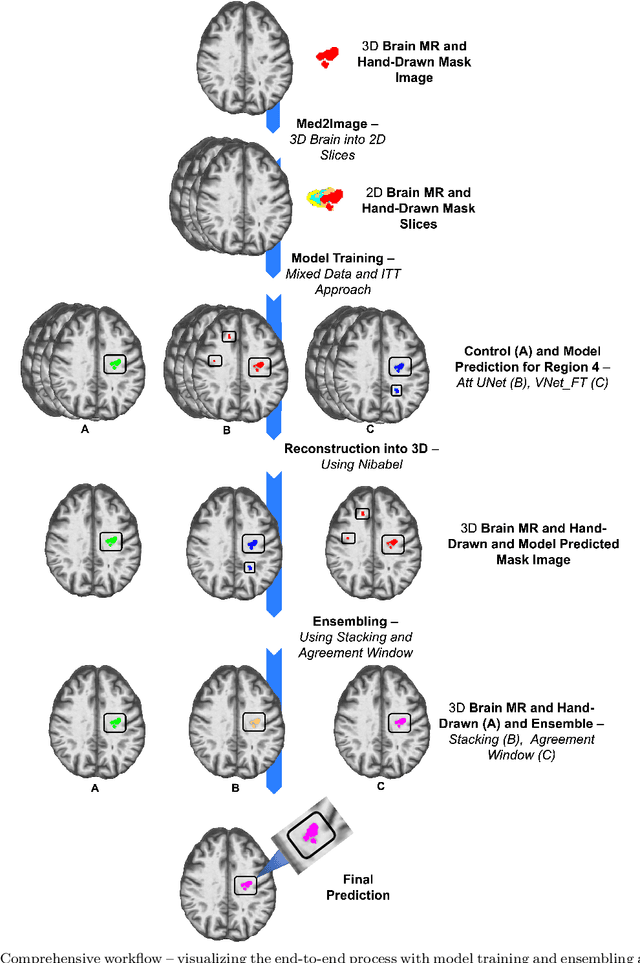
Abstract:A major challenge in stroke research and stroke recovery predictions is the determination of a stroke lesion's extent and its impact on relevant brain systems. Manual segmentation of stroke lesions from 3D magnetic resonance (MR) imaging volumes, the current gold standard, is not only very time-consuming, but its accuracy highly depends on the operator's experience. As a result, there is a need for a fully automated segmentation method that can efficiently and objectively measure lesion extent and the impact of each lesion to predict impairment and recovery potential which might be beneficial for clinical, translational, and research settings. We have implemented and tested a fully automatic method for stroke lesion segmentation which was developed using eight different 2D-model architectures trained via transfer learning (TL) and mixed data approaches. Additionally, the final prediction was made using a novel ensemble method involving stacking and agreement window. Our novel method was evaluated in a novel in-house dataset containing 22 T1w brain MR images, which were challenging in various perspectives, but mostly because they included T1w MR images from the subacute (which typically less well defined T1 lesions) and chronic stroke phase (which typically means well defined T1-lesions). Cross-validation results indicate that our new method can efficiently and automatically segment lesions fast and with high accuracy compared to ground truth. In addition to segmentation, we provide lesion volume and weighted lesion load of relevant brain systems based on the lesions' overlap with a canonical structural motor system that stretches from the cortical motor region to the lowest end of the brain stem.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge