Ahsan Raza Siyal
Schrödinger-Inspired Time-Evolution for 4D Deformation Forecasting
Jan 31, 2026Abstract:Spatiotemporal forecasting of complex three-dimensional phenomena (4D: 3D + time) is fundamental to applications in medical imaging, fluid and material dynamics, and geophysics. In contrast to unconstrained neural forecasting models, we propose a Schrödinger-inspired, physics-guided neural architecture that embeds an explicit time-evolution operator within a deep convolutional framework for 4D prediction. From observed volumetric sequences, the model learns voxelwise amplitude, phase, and potential fields that define a complex-valued wavefunction $ψ= A e^{iφ}$, which is evolved forward in time using a differentiable, unrolled Schrödinger time stepper. This physics-guided formulation yields several key advantages: (i) temporal stability arising from the structured evolution operator, which mitigates drift and error accumulation in long-horizon forecasting; (ii) an interpretable latent representation, where phase encodes transport dynamics, amplitude captures structural intensity, and the learned potential governs spatiotemporal interactions; and (iii) natural compatibility with deformation-based synthesis, which is critical for preserving anatomical fidelity in medical imaging applications. By integrating physical priors directly into the learning process, the proposed approach combines the expressivity of deep networks with the robustness and interpretability of physics-based modeling. We demonstrate accurate and stable prediction of future 4D states, including volumetric intensities and deformation fields, on synthetic benchmarks that emulate realistic shape deformations and topological changes. To our knowledge, this is the first end-to-end 4D neural forecasting framework to incorporate a Schrödinger-type evolution operator, offering a principled pathway toward interpretable, stable, and anatomically consistent spatiotemporal prediction.
A lightweight residual network for unsupervised deformable image registration
Jun 14, 2024



Abstract:Accurate volumetric image registration is highly relevant for clinical routines and computer-aided medical diagnosis. Recently, researchers have begun to use transformers in learning-based methods for medical image registration, and have achieved remarkable success. Due to the strong global modeling capability, Transformers are considered a better option than convolutional neural networks (CNNs) for registration. However, they use bulky models with huge parameter sets, which require high computation edge devices for deployment as portable devices or in hospitals. Transformers also need a large amount of training data to produce significant results, and it is often challenging to collect suitable annotated data. Although existing CNN-based image registration can offer rich local information, their global modeling capability is poor for handling long-distance information interaction and limits registration performance. In this work, we propose a CNN-based registration method with an enhanced receptive field, a low number of parameters, and significant results on a limited training dataset. For this, we propose a residual U-Net with embedded parallel dilated-convolutional blocks to enhance the receptive field. The proposed method is evaluated on inter-patient and atlas-based datasets. We show that the performance of the proposed method is comparable and slightly better than transformer-based methods by using only $\SI{1.5}{\percent}$ of its number of parameters.
Unsupervised Joint Image Transfer and Uncertainty Quantification using Patch Invariant Networks
Jul 09, 2022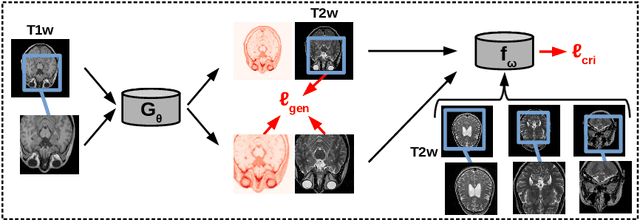
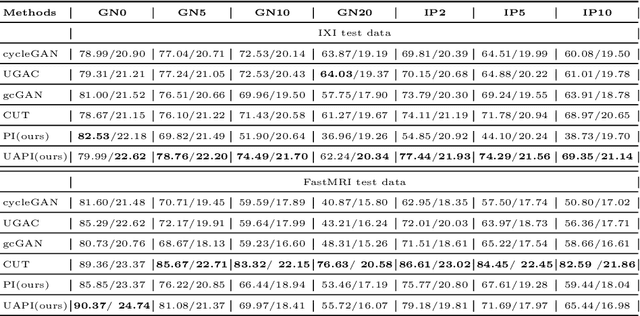

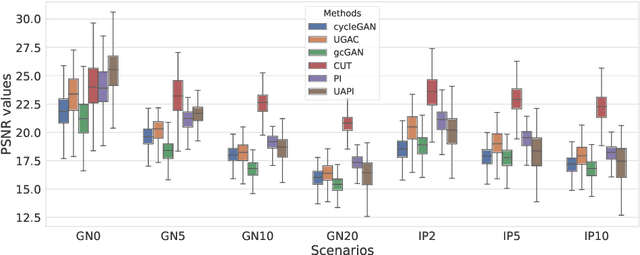
Abstract:Unsupervised image transfer enables intra- and inter-modality transfer for medical applications where a large amount of paired training data is not abundant. To ensure a structure-preserving mapping from the input to the target domain, existing methods for unpaired medical image transfer are commonly based on cycle-consistency, causing additional computation resources and instability due to the learning of an inverse mapping. This paper presents a novel method for uni-directional domain mapping where no paired data is needed throughout the entire training process. A reasonable transfer is ensured by employing the GAN architecture and a novel generator loss based on patch invariance. To be more precise, generator outputs are evaluated and compared on different scales, which brings increased attention to high-frequency details as well as implicit data augmentation. This novel term also gives the opportunity to predict aleatoric uncertainty by modeling an input-dependent scale map for the patch residuals. The proposed method is comprehensively evaluated on three renowned medical databases. Superior accuracy on these datasets compared to four different state-of-the-art methods for unpaired image transfer suggests the great potential of this approach for uncertainty-aware medical image translation. Implementation of the proposed framework is released here: https://github.com/anger-man/unsupervised-image-transfer-and-uq.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge