Ahmad F. Al Musawi
A Review of Link Prediction Applications in Network Biology
Dec 03, 2023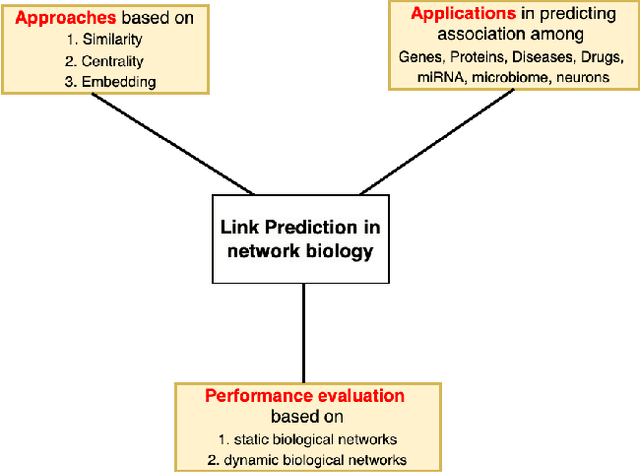
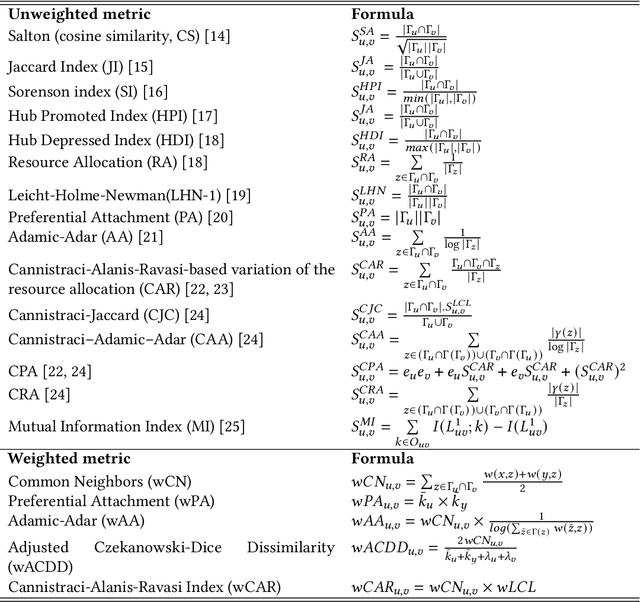
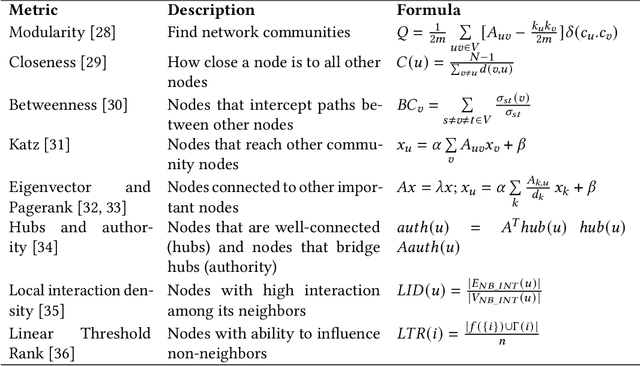


Abstract:In the domain of network biology, the interactions among heterogeneous genomic and molecular entities are represented through networks. Link prediction (LP) methodologies are instrumental in inferring missing or prospective associations within these biological networks. In this review, we systematically dissect the attributes of local, centrality, and embedding-based LP approaches, applied to static and dynamic biological networks. We undertake an examination of the current applications of LP metrics for predicting links between diseases, genes, proteins, RNA, microbiomes, drugs, and neurons. We carry out comprehensive performance evaluations on established biological network datasets to show the practical applications of standard LP models. Moreover, we compare the similarity in prediction trends among the models and the specific network attributes that contribute to effective link prediction, before underscoring the role of LP in addressing the formidable challenges prevalent in biological systems, ranging from noise, bias, and data sparseness to interpretability. We conclude the review with an exploration of the essential characteristics expected from future LP models, poised to advance our comprehension of the intricate interactions governing biological systems.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge