Variational inference for rare variant detection in deep, heterogeneous next-generation sequencing data
Paper and Code
Apr 22, 2016
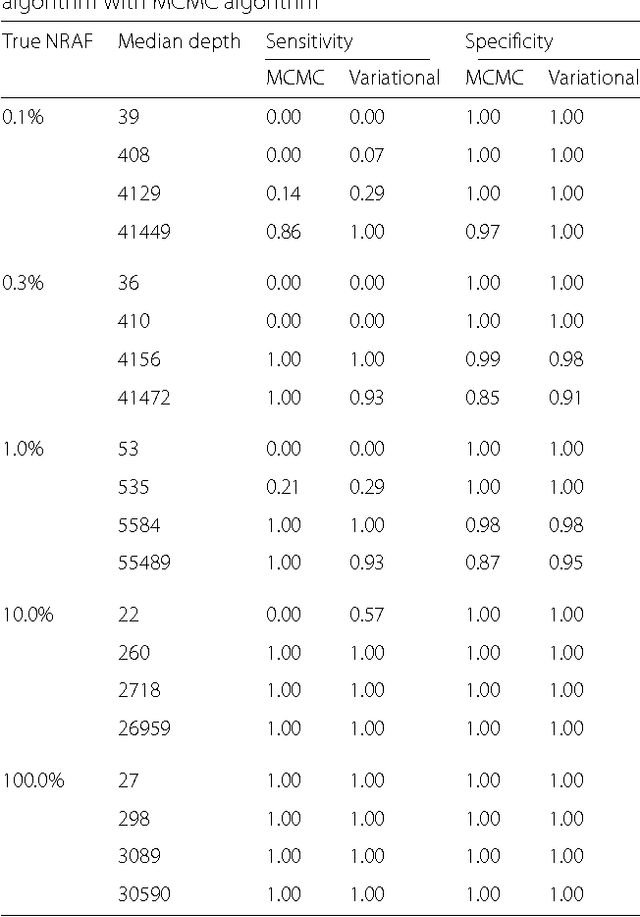
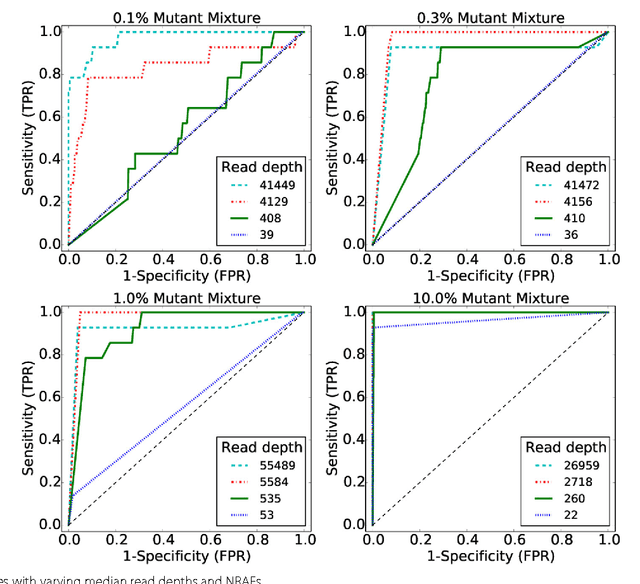

The detection of rare variants is important for understanding the genetic heterogeneity in mixed samples. Recently, next-generation sequencing (NGS) technologies have enabled the identification of single nucleotide variants (SNVs) in mixed samples with high resolution. Yet, the noise inherent in the biological processes involved in next-generation sequencing necessitates the use of statistical methods to identify true rare variants. We propose a novel Bayesian statistical model and a variational expectation-maximization (EM) algorithm to estimate non-reference allele frequency (NRAF) and identify SNVs in heterogeneous cell populations. We demonstrate that our variational EM algorithm has comparable sensitivity and specificity compared with a Markov Chain Monte Carlo (MCMC) sampling inference algorithm, and is more computationally efficient on tests of low coverage ($27\times$ and $298\times$) data. Furthermore, we show that our model with a variational EM inference algorithm has higher specificity than many state-of-the-art algorithms. In an analysis of a directed evolution longitudinal yeast data set, we are able to identify a time-series trend in non-reference allele frequency and detect novel variants that have not yet been reported. Our model also detects the emergence of a beneficial variant earlier than was previously shown, and a pair of concomitant variants.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge