Unsupervised Detection of Lesions in Brain MRI using constrained adversarial auto-encoders
Paper and Code
Jun 13, 2018

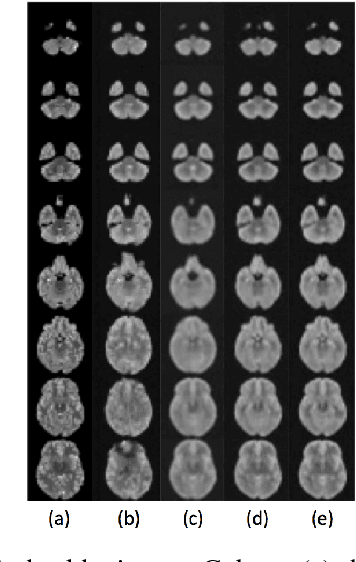
Lesion detection in brain Magnetic Resonance Images (MRI) remains a challenging task. State-of-the-art approaches are mostly based on supervised learning making use of large annotated datasets. Human beings, on the other hand, even non-experts, can detect most abnormal lesions after seeing a handful of healthy brain images. Replicating this capability of using prior information on the appearance of healthy brain structure to detect lesions can help computers achieve human level abnormality detection, specifically reducing the need for numerous labeled examples and bettering generalization of previously unseen lesions. To this end, we study detection of lesion regions in an unsupervised manner by learning data distribution of brain MRI of healthy subjects using auto-encoder based methods. We hypothesize that one of the main limitations of the current models is the lack of consistency in latent representation. We propose a simple yet effective constraint that helps mapping of an image bearing lesion close to its corresponding healthy image in the latent space. We use the Human Connectome Project dataset to learn distribution of healthy-appearing brain MRI and report improved detection, in terms of AUC, of the lesions in the BRATS challenge dataset.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge