Ultrasound segmentation using U-Net: learning from simulated data and testing on real data
Paper and Code
Apr 24, 2019
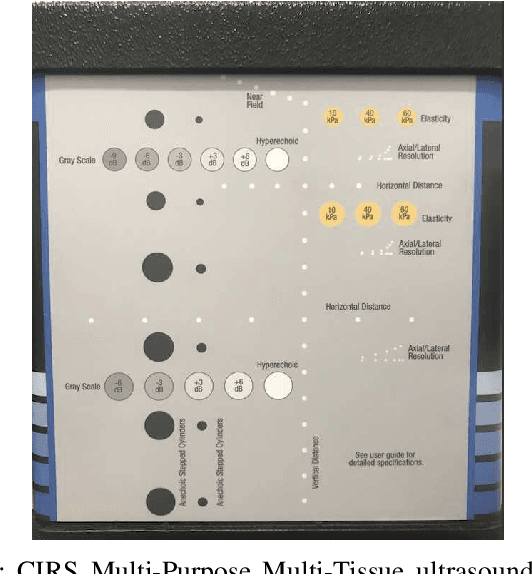
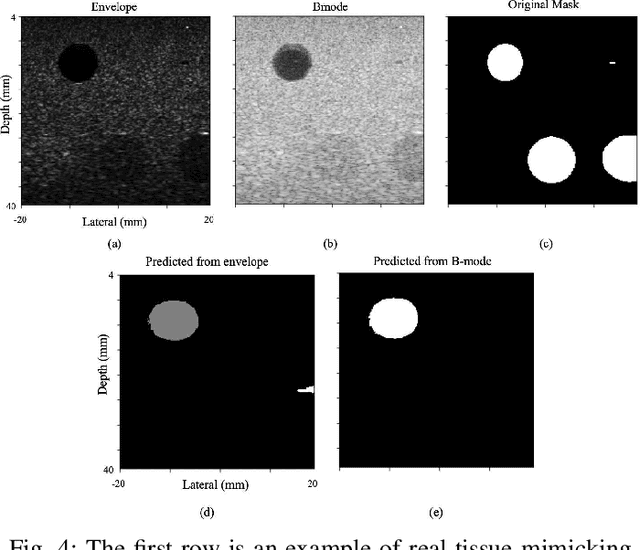

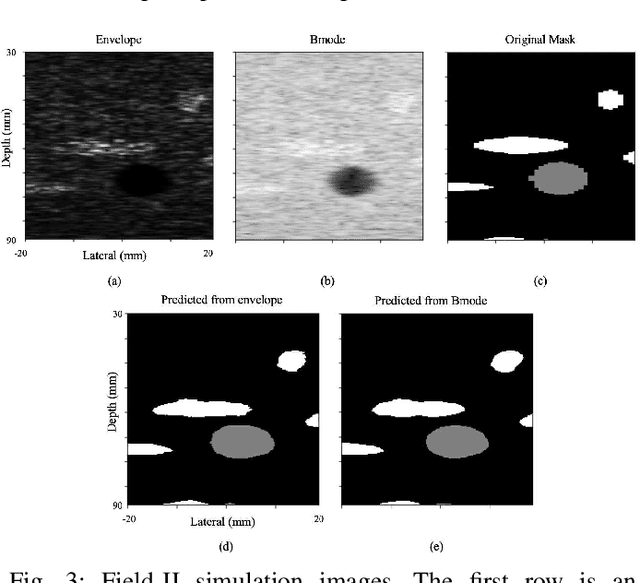

Segmentation of ultrasound images is an essential task in both diagnosis and image-guided interventions given the ease-of-use and low cost of this imaging modality. As manual segmentation is tedious and time consuming, a growing body of research has focused on the development of automatic segmentation algorithms. Deep learning algorithms have shown remarkable achievements in this regard; however, they need large training datasets. Unfortunately, preparing large labeled datasets in ultrasound images is prohibitively difficult. Therefore, in this study, we propose the use of simulated ultrasound (US) images for training the U-Net deep learning segmentation architecture and test on tissue-mimicking phantom data collected by an ultrasound machine. We demonstrate that the trained architecture on the simulated data is transferrable to real data, and therefore, simulated data can be considered as an alternative training dataset when real datasets are not available. The second contribution of this paper is that we train our U- Net network on envelope and B-mode images of the simulated dataset, and test the trained network on real envelope and B- mode images of phantom, respectively. We show that test results are superior for the envelope data compared to B-mode image.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge