Transcriptome-supervised classification of tissue morphology using deep learning
Paper and Code
Dec 07, 2023
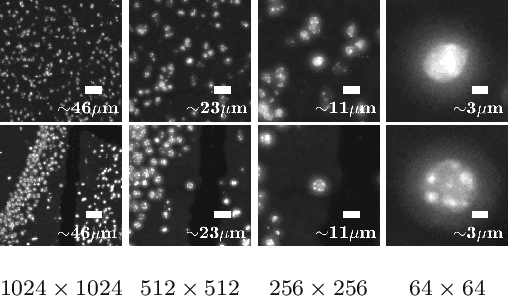

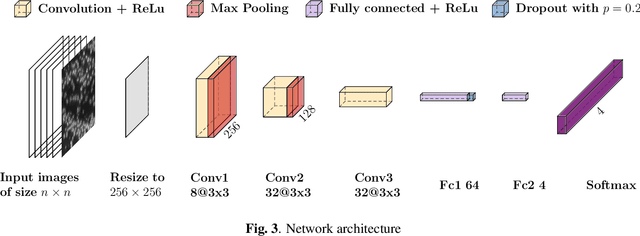
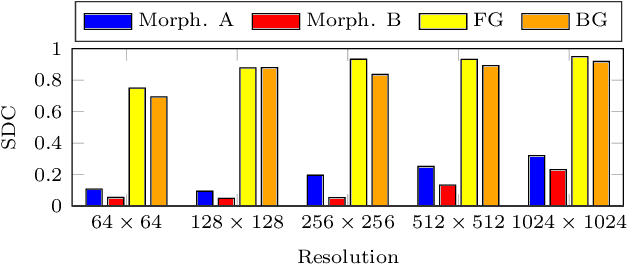
Deep learning has proven to successfully learn variations in tissue and cell morphology. Training of such models typically relies on expensive manual annotations. Here we conjecture that spatially resolved gene expression, e.i., the transcriptome, can be used as an alternative to manual annotations. In particular, we trained five convolutional neural networks with patches of different size extracted from locations defined by spatially resolved gene expression. The network is trained to classify tissue morphology related to two different genes, general tissue, as well as background, on an image of fluorescence stained nuclei in a mouse brain coronal section. Performance is evaluated on an independent tissue section from a different mouse brain, reaching an average Dice score of 0.51. Results may indicate that novel techniques for spatially resolved transcriptomics together with deep learning may provide a unique and unbiased way to find genotype-phenotype relationships.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge