The Many-Body Expansion Combined with Neural Networks
Paper and Code
Sep 22, 2016
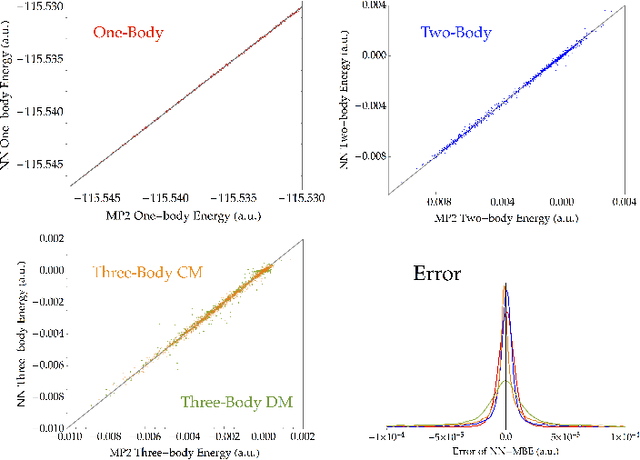
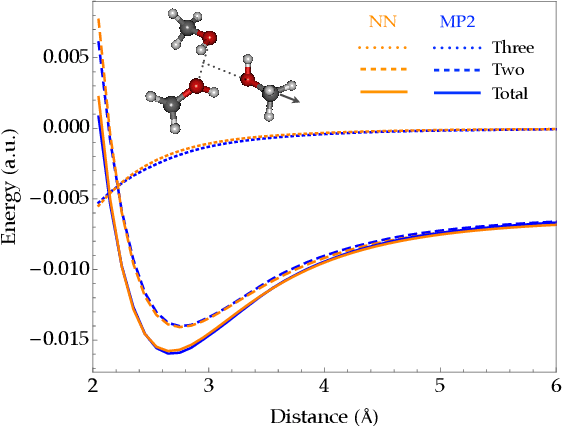
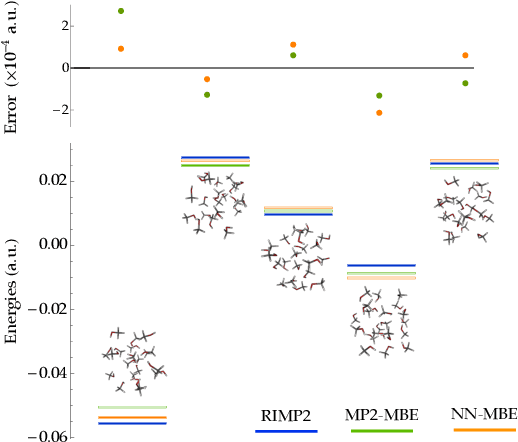
Fragmentation methods such as the many-body expansion (MBE) are a common strategy to model large systems by partitioning energies into a hierarchy of decreasingly significant contributions. The number of fragments required for chemical accuracy is still prohibitively expensive for ab-initio MBE to compete with force field approximations for applications beyond single-point energies. Alongside the MBE, empirical models of ab-initio potential energy surfaces have improved, especially non-linear models based on neural networks (NN) which can reproduce ab-initio potential energy surfaces rapidly and accurately. Although they are fast, NNs suffer from their own curse of dimensionality; they must be trained on a representative sample of chemical space. In this paper we examine the synergy of the MBE and NN's, and explore their complementarity. The MBE offers a systematic way to treat systems of arbitrary size and intelligently sample chemical space. NN's reduce, by a factor in excess of $10^6$ the computational overhead of the MBE and reproduce the accuracy of ab-initio calculations without specialized force fields. We show they are remarkably general, providing comparable accuracy with drastically different chemical embeddings. To assess this we test a new chemical embedding which can be inverted to predict molecules with desired properties.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge