Stable deep neural network architectures for mitochondria segmentation on electron microscopy volumes
Paper and Code
Apr 08, 2021
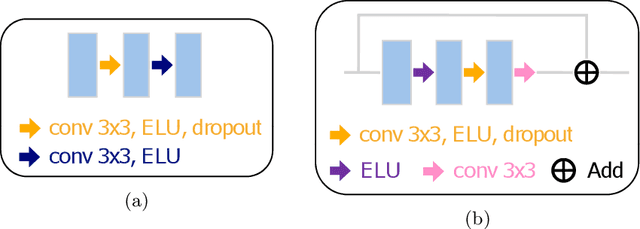

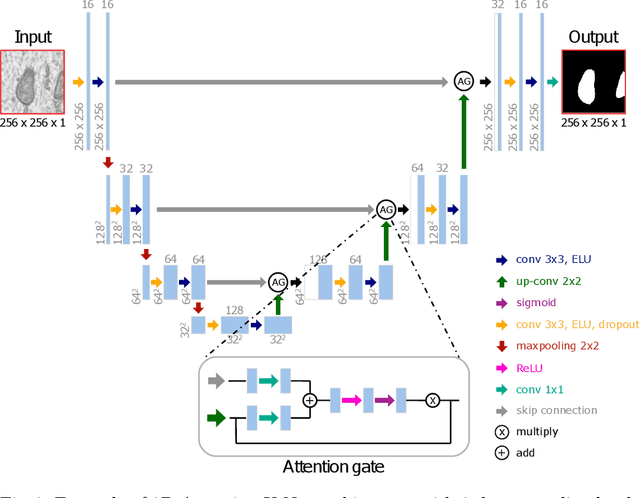
Electron microscopy (EM) allows the identification of intracellular organelles such as mitochondria, providing insights for clinical and scientific studies. In recent years, a number of novel deep learning architectures have been published reporting superior performance, or even human-level accuracy, compared to previous approaches on public mitochondria segmentation datasets. Unfortunately, many of these publications do not make neither the code nor the full training details public to support the results obtained, leading to reproducibility issues and dubious model comparisons. For that reason, and following a recent code of best practices for reporting experimental results, we present an extensive study of the state-of-the-art deep learning architectures for the segmentation of mitochondria on EM volumes, and evaluate the impact in performance of different variations of 2D and 3D U-Net-like models for this task. To better understand the contribution of each component, a common set of pre- and post-processing operations has been implemented and tested with each approach. Moreover, an exhaustive sweep of hyperparameters values for all architectures have been performed and each configuration has been run multiple times to report the mean and standard deviation values of the evaluation metrics. Using this methodology, we found very stable architectures and hyperparameter configurations that consistently obtain state-of-the-art results in the well-known EPFL Hippocampus mitochondria segmentation dataset. Furthermore, we have benchmarked our proposed models on two other available datasets, Lucchi++ and Kasthuri++, where they outperform all previous works. The code derived from this research and its documentation are publicly available.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge