Sparse Hierachical Extrapolated Parametric Methods for Cortical Data Analysis
Paper and Code
Apr 27, 2017
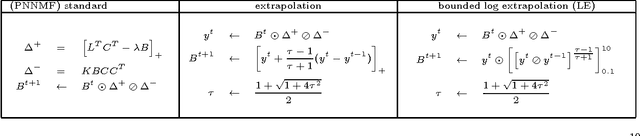
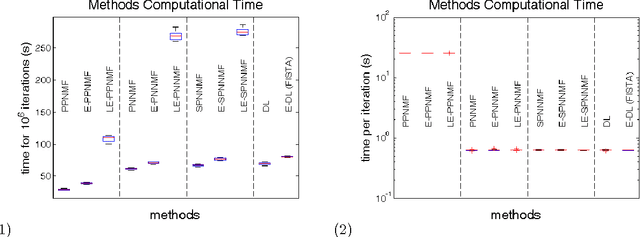
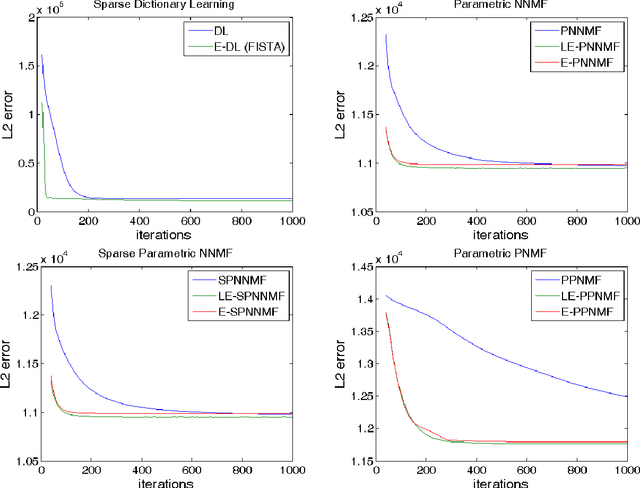
Many neuroimaging studies focus on the cortex, in order to benefit from better signal to noise ratios and reduced computational burden. Cortical data are usually projected onto a reference mesh, where subsequent analyses are carried out. Several multiscale approaches have been proposed for analyzing these surface data, such as spherical harmonics and graph wavelets. As far as we know, however, the hierarchical structure of the template icosahedral meshes used by most neuroimaging software has never been exploited for cortical data factorization. In this paper, we demonstrate how the structure of the ubiquitous icosahedral meshes can be exploited by data factorization methods such as sparse dictionary learning, and we assess the optimization speed-up offered by extrapolation methods in this context. By testing different sparsity-inducing norms, extrapolation methods, and factorization schemes, we compare the performances of eleven methods for analyzing four datasets: two structural and two functional MRI datasets obtained by processing the data publicly available for the hundred unrelated subjects of the Human Connectome Project. Our results demonstrate that, depending on the level of details requested, a speedup of several orders of magnitudes can be obtained.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge