SlideGCD: Slide-based Graph Collaborative Training with Knowledge Distillation for Whole Slide Image Classification
Paper and Code
Jul 12, 2024
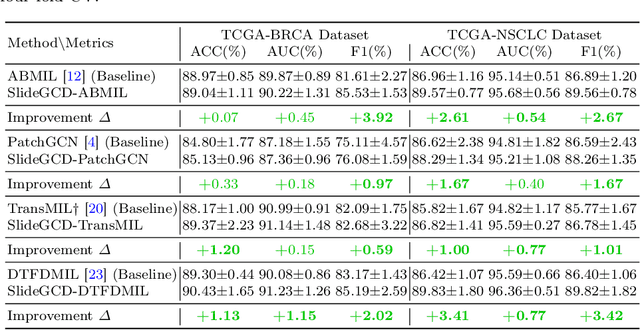
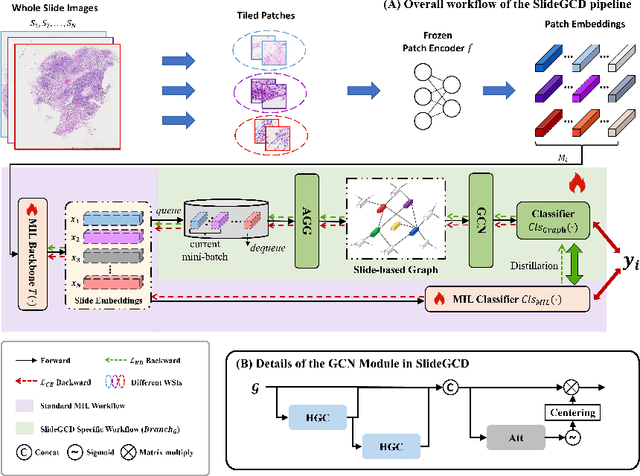
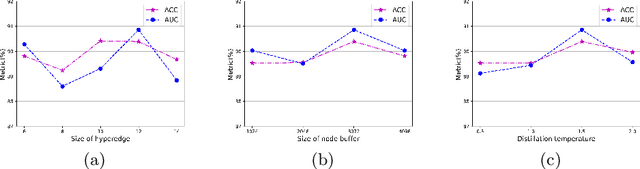
Existing WSI analysis methods lie on the consensus that histopathological characteristics of tumors are significant guidance for cancer diagnostics. Particularly, as the evolution of cancers is a continuous process, the correlations and differences across various stages, anatomical locations and patients should be taken into account. However, recent research mainly focuses on the inner-contextual information in a single WSI, ignoring the correlations between slides. To verify whether introducing the slide inter-correlations can bring improvements to WSI representation learning, we propose a generic WSI analysis pipeline SlideGCD that considers the existing multi-instance learning (MIL) methods as the backbone and forge the WSI classification task as a node classification problem. More specifically, SlideGCD declares a node buffer that stores previous slide embeddings for subsequent extensive slide-based graph construction and conducts graph learning to explore the inter-correlations implied in the slide-based graph. Moreover, we frame the MIL classifier and graph learning into two parallel workflows and deploy the knowledge distillation to transfer the differentiable information to the graph neural network. The consistent performance boosting, brought by SlideGCD, of four previous state-of-the-art MIL methods is observed on two TCGA benchmark datasets. The code is available at https://github.com/HFUT-miaLab/SlideGCD.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge