Simultaneous Multiple Surface Segmentation Using Deep Learning
Paper and Code
May 19, 2017


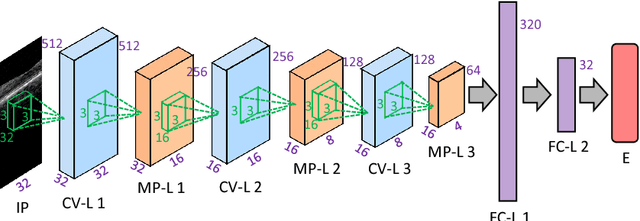
The task of automatically segmenting 3-D surfaces representing boundaries of objects is important for quantitative analysis of volumetric images, and plays a vital role in biomedical image analysis. Recently, graph-based methods with a global optimization property have been developed and optimized for various medical imaging applications. Despite their widespread use, these require human experts to design transformations, image features, surface smoothness priors, and re-design for a different tissue, organ or imaging modality. Here, we propose a Deep Learning based approach for segmentation of the surfaces in volumetric medical images, by learning the essential features and transformations from training data, without any human expert intervention. We employ a regional approach to learn the local surface profiles. The proposed approach was evaluated on simultaneous intraretinal layer segmentation of optical coherence tomography (OCT) images of normal retinas and retinas affected by age related macular degeneration (AMD). The proposed approach was validated on 40 retina OCT volumes including 20 normal and 20 AMD subjects. The experiments showed statistically significant improvement in accuracy for our approach compared to state-of-the-art graph based optimal surface segmentation with convex priors (G-OSC). A single Convolution Neural Network (CNN) was used to learn the surfaces for both normal and diseased images. The mean unsigned surface positioning errors obtained by G-OSC method 2.31 voxels (95% CI 2.02-2.60 voxels) was improved to $1.27$ voxels (95% CI 1.14-1.40 voxels) using our new approach. On average, our approach takes 94.34 s, requiring 95.35 MB memory, which is much faster than the 2837.46 s and 6.87 GB memory required by the G-OSC method on the same computer system.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge