Scalable Knowledge Graph Construction and Inference on Human Genome Variants
Paper and Code
Dec 07, 2023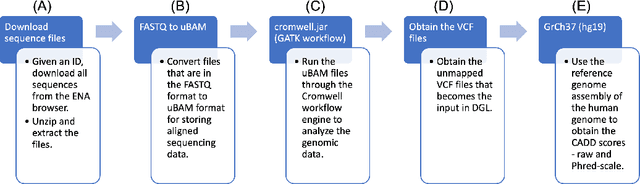
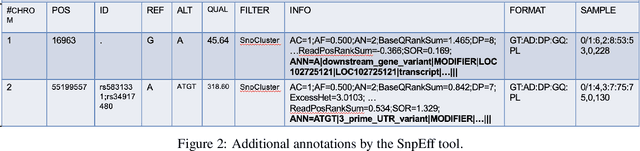



Real-world knowledge can be represented as a graph consisting of entities and relationships between the entities. The need for efficient and scalable solutions arises when dealing with vast genomic data, like RNA-sequencing. Knowledge graphs offer a powerful approach for various tasks in such large-scale genomic data, such as analysis and inference. In this work, variant-level information extracted from the RNA-sequences of vaccine-na\"ive COVID-19 patients have been represented as a unified, large knowledge graph. Variant call format (VCF) files containing the variant-level information were annotated to include further information for each variant. The data records in the annotated files were then converted to Resource Description Framework (RDF) triples. Each VCF file obtained had an associated CADD scores file that contained the raw and Phred-scaled scores for each variant. An ontology was defined for the VCF and CADD scores files. Using this ontology and the extracted information, a large, scalable knowledge graph was created. Available graph storage was then leveraged to query and create datasets for further downstream tasks. We also present a case study using the knowledge graph and perform a classification task using graph machine learning. We also draw comparisons between different Graph Neural Networks (GNNs) for the case study.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge