Scalable Fragment-Based 3D Molecular Design with Reinforcement Learning
Paper and Code
Feb 01, 2022
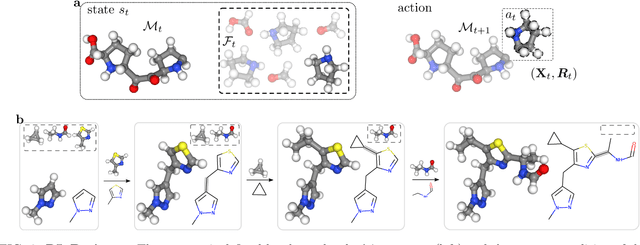
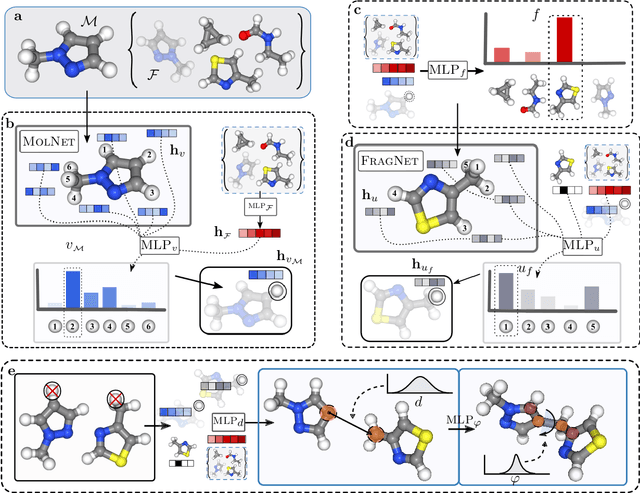
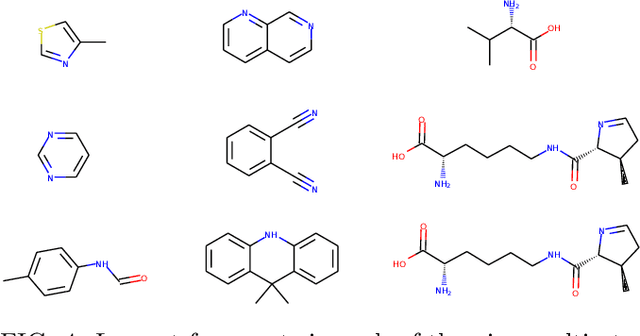
Machine learning has the potential to automate molecular design and drastically accelerate the discovery of new functional compounds. Towards this goal, generative models and reinforcement learning (RL) using string and graph representations have been successfully used to search for novel molecules. However, these approaches are limited since their representations ignore the three-dimensional (3D) structure of molecules. In fact, geometry plays an important role in many applications in inverse molecular design, especially in drug discovery. Thus, it is important to build models that can generate molecular structures in 3D space based on property-oriented geometric constraints. To address this, one approach is to generate molecules as 3D point clouds by sequentially placing atoms at locations in space -- this allows the process to be guided by physical quantities such as energy or other properties. However, this approach is inefficient as placing individual atoms makes the exploration unnecessarily deep, limiting the complexity of molecules that can be generated. Moreover, when optimizing a molecule, organic and medicinal chemists use known fragments and functional groups, not single atoms. We introduce a novel RL framework for scalable 3D design that uses a hierarchical agent to build molecules by placing molecular substructures sequentially in 3D space, thus attempting to build on the existing human knowledge in the field of molecular design. In a variety of experiments with different substructures, we show that our agent, guided only by energy considerations, can efficiently learn to produce molecules with over 100 atoms from many distributions including drug-like molecules, organic LED molecules, and biomolecules.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge