ProtoKD: Learning from Extremely Scarce Data for Parasite Ova Recognition
Paper and Code
Sep 18, 2023
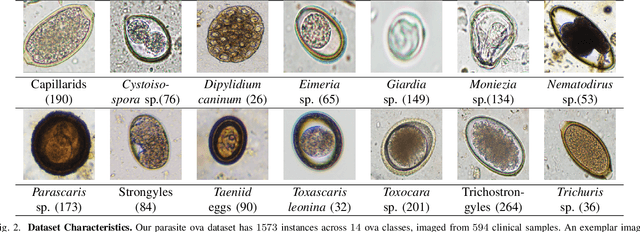
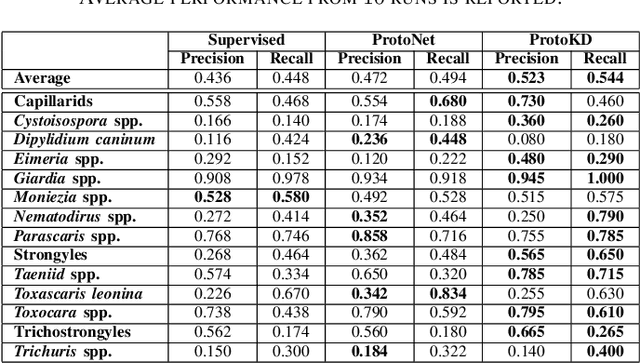
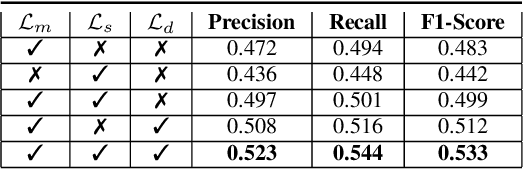
Developing reliable computational frameworks for early parasite detection, particularly at the ova (or egg) stage is crucial for advancing healthcare and effectively managing potential public health crises. While deep learning has significantly assisted human workers in various tasks, its application and diagnostics has been constrained by the need for extensive datasets. The ability to learn from an extremely scarce training dataset, i.e., when fewer than 5 examples per class are present, is essential for scaling deep learning models in biomedical applications where large-scale data collection and annotation can be expensive or not possible (in case of novel or unknown infectious agents). In this study, we introduce ProtoKD, one of the first approaches to tackle the problem of multi-class parasitic ova recognition using extremely scarce data. Combining the principles of prototypical networks and self-distillation, we can learn robust representations from only one sample per class. Furthermore, we establish a new benchmark to drive research in this critical direction and validate that the proposed ProtoKD framework achieves state-of-the-art performance. Additionally, we evaluate the framework's generalizability to other downstream tasks by assessing its performance on a large-scale taxonomic profiling task based on metagenomes sequenced from real-world clinical data.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge