Programming and Training Rate-Independent Chemical Reaction Networks
Paper and Code
Sep 20, 2021
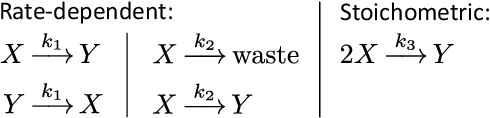
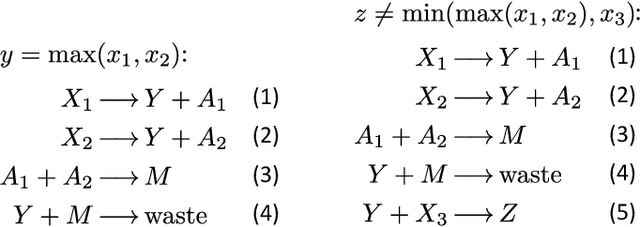
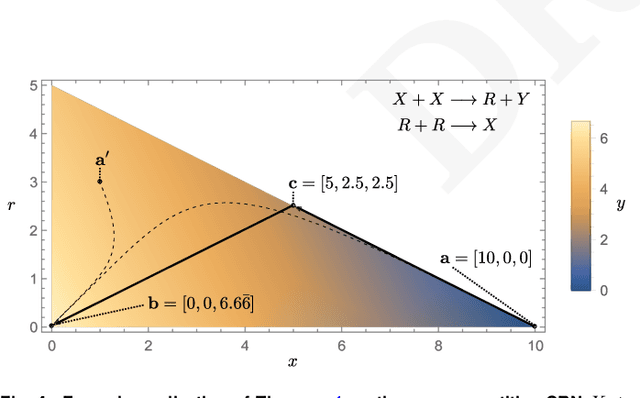
Embedding computation in biochemical environments incompatible with traditional electronics is expected to have wide-ranging impact in synthetic biology, medicine, nanofabrication and other fields. Natural biochemical systems are typically modeled by chemical reaction networks (CRNs), and CRNs can be used as a specification language for synthetic chemical computation. In this paper, we identify a class of CRNs called non-competitive (NC) whose equilibria are absolutely robust to reaction rates and kinetic rate law, because their behavior is captured solely by their stoichiometric structure. Unlike prior work on rate-independent CRNs, checking non-competition and using it as a design criterion is easy and promises robust output. We also present a technique to program NC-CRNs using well-founded deep learning methods, showing a translation procedure from rectified linear unit (ReLU) neural networks to NC-CRNs. In the case of binary weight ReLU networks, our translation procedure is surprisingly tight in the sense that a single bimolecular reaction corresponds to a single ReLU node and vice versa. This compactness argues that neural networks may be a fitting paradigm for programming rate-independent chemical computation. As proof of principle, we demonstrate our scheme with numerical simulations of CRNs translated from neural networks trained on traditional machine learning datasets (IRIS and MNIST), as well as tasks better aligned with potential biological applications including virus detection and spatial pattern formation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge